ABSTRACT
Cruciferous vegetables contain glucosinolates that, after conversion to isothiocyanates (ITC), are capable of inducing cytoprotective genes. We examined whether broccoli seeds can elicit a chemoprotective response in mouse organs and rodent cell lines and investigated whether this response requires nuclear factor-erythroid 2 p45-related factor 2 (Nrf2). The seeds studied contained glucosinolate at 40 mmol/kg, of which 59% comprised glucoiberin, 19% sinigrin, 8% glucoraphanin, and 7% progoitrin. Dietary administration of broccoli seeds to nrf2+/+ and nrf2−/− mice produced a ∼1.5-fold increase in NAD(P)H:quinone oxidoreductase 1 (NQO1) and glutathione S-transferase (GST) activities in stomach, small intestine, and liver of wild-type mice but not in mutant mice; increased transferase activity was associated with elevated levels of GSTA1/2, GSTA3, and GSTM1/2 subunits. These seeds also increased significantly the level of glutamate cysteine ligase catalytic (GCLC) subunit in the stomach and the small intestine of nrf2+/+ mice but not nrf2−/− mice. An aqueous broccoli seed extract was prepared for treatment of cultured cells that contained ITC at ∼600 μmol/L, composed of 61% 3-methylsulfinylpropyl ITC, 30% sulforaphane, 4% allyl ITC, and 4% 3-butenyl ITC. This extract induced GSTA1/2, GSTA3, NQO1, and GCLC between 3-fold and 10-fold in mouse Hepa-1c1c7 and rat liver RL-34 cells. The broccoli seed extract affected increases in GSTA3, GSTM1, and NQO1 proteins in nrf2+/+ mouse embryonic fibroblasts but not in nrf2−/− mouse embryonic fibroblasts. These experiments show that broccoli seeds are effective at inducing antioxidant and detoxication proteins, both in vivo and ex vivo, in an Nrf2-dependent manner.
Glutathione S-transferase M3 (brain), also known as GSTM3, is an enzyme which in humans is encoded by the GSTM3 gene. Glutathione S-transferase Mu 4 is an enzyme that in humans is encoded by the GSTM4 gene.Cytosolic and membrane-bound forms of glutathione S-transferase are encoded by two distinct supergene families. A rat testicular Mu-class glutathioneS-transferase (GST) resolved by reversed-phase high performance liquid chromatography cross-reacted with peptide sequence-specific antisera raised against the human hGSTM3 subunit. Electrospray ionization mass spectrometry indicated that this rat GST subunit (designated rGSTM5 in this report) has a significantly greater molecular mass (26,541 Da) than the other rat GST subunits.
antioxidant response element, chemoprevention, glucosinolates, sulforaphane, myrosinase
Individuals who have a high dietary intake of fruit and vegetables appear to have a lower risk of cancer (1). Among vegetables with anticarcinogenic properties, members of the Cruciferae family have been reported to protect against neoplastic disease at a variety of sites, such as the gastrointestinal tract and the lungs (2–6).
The cancer chemopreventive effect of cruciferous vegetables has been attributed to the fact that they contain high levels of glucosinolates (7,8). During food preparation and eating, these glucosinolates are hydrolyzed by the plant enzyme myrosinase to yield a complex number of breakdown products, including isothiocyanates (ITC),4 thiocyanates, cyanides, nitriles, and epithio-containing compounds (7–9). Some of these breakdown products, and, in particular, ITCs can increase the levels of detoxication enzymes in rodent organs and in mouse, rat, and human cell lines (10–17). Inducible proteins include the drug-metabolizing enzymes aldo-keto reductase, NAD(P)H:quinone oxidoreductase 1 (NQO1), and glutathione S-transferase (GST). Increases in the levels of these detoxication enzymes would be expected to confer protection against chemical carcinogens such as benzo[a]pyrene, and, in experimental models, this prediction appears to hold true (14,18). Less well appreciated is the fact that glucosinolate breakdown products also induce antioxidant proteins, such as the glutamate cysteine ligase catalytic (GCLC) and glutamate cysteine ligase modifier (GCLM) subunits, that catalyze the rate-limiting step in the formation of reduced glutathione (19,20). They also induce glutathione reductase, ferritin, and glucose-6-phosphate dehydrogenase (20). Increases in the levels of detoxication enzymes and antioxidant proteins would be expected to protect against reactive oxygen species and the harmful metabolites they generate as a consequence of damaging cellular membranes, proteins, and nucleic acids (21).
Many genes encoding detoxication and antioxidant proteins are regulated by nuclear factor-erythroid 2 p45-related factor 2 (Nrf2) (22). This basic-region leucine zipper transcription factor mediates the transcriptional activation of genes in response to oxidative and electrophile stress. Under normal homeostatic conditions, Nrf2 protein has a short half-life, being targeted for proteasomal degradation by Keap1 (23–26). Oxidants and electrophiles interfere with Keap1-facilitated degradation of Nrf2, causing it to become more stable. This process involves oxidation, modification, or both, of cysteine residues 273 and 288 in Keap1 by the inducing compounds (27). Induction of NQO1, GST, GCLC, and GCLM genes by Nrf2 occurs through it being recruited to antioxidant response elements (ARE) in their gene promoters (28); Nrf2 binds the ARE as a heterodimer with small Maf proteins (29,30). Mice in which the nrf2 gene has been disrupted by targeted homologous recombination have lower constitutive levels of NQO1 and GST proteins in liver and small intestine (19,31,32). Furthermore, nrf2−/− mice are either unable to respond or have a blunted response to the model cancer chemopreventive agent butylated hydroxyanisole (19,33,34).
Most investigations into the ability of plant chemicals to increase antioxidant gene expression used highly purified compounds as inducing agents (11–20). Thus, ITCs such as sulforaphane have been shown to increase NQO1 enzyme activity in the mouse liver Hepa-1c1c7 cell line (35). Frequently, it is unclear whether the concentration of phytochemical used in cell culture experiments is physiologically relevant and whether, because of limitations caused by bioavailability or disposition, the dose of chemical used can be achieved in target tissues in vivo. The question of whether extracts of cruciferous plants are as effective as purified phytochemicals at stimulating gene expression is seldom addressed.
In this study, we investigated whether broccoli seeds, either in the diet or as aqueous extracts, can affect induction of antioxidant and detoxication genes in vivo, in transformed cells, and in nontransformed cells. We also tested the hypothesis that Nrf2, through stimulating ARE-driven gene transcription, is essential for gene induction by broccoli-derived phytochemicals.
Materials and methods
Chemicals
Allyl ITC (AITC) and sulforaphane were obtained from Aldrich and LKT Laboratories, respectively. All other chemicals used were of the highest purity that was available from commercial suppliers.
Broccoli seeds
Broccoli seeds were purchased from Thompson and Morgan.
Processing of broccoli seeds for induction experiments
The broccoli seeds were processed at room temperature (20°C). Extracts were prepared by crushing 10 g seeds (dry weight), by pestle and mortar, to a fine powder. For mice feeding experiments, crushed broccoli seeds were added directly to powdered RM1 laboratory animal feed (SDS) at 15% by weight. For cell culture experiments, the broccoli seed powder was suspended in 3 volumes of distilled water and was mixed vigorously for 5 min. The suspension was centrifuged at 800 × g for 10 min before being filtered through a 0.2-μm sterile filter. Aliquots (1 mL) of the aqueous filtered extract were snap-frozen in liquid nitrogen and were stored at −70°C before use; the entire process from crushing the broccoli seeds to snap-freezing the filtered aqueous extract was completed within 30 min. The frozen extracts were thawed rapidly and diluted 1/1000 in 6 mL of medium for cell culture experiments that were conducted in 60-mm dishes.
Analysis of glucosinolates and ITCs in broccoli seeds
Glucosinolates and corresponding ITCs were identified by liquid chromatography with triple quadrupole MS detection (LC-MS/MS). Standard reference glucosinolates were isolated and purified from Brassica seeds by modification of published methods (36), and the related ITCs were prepared by myrosinase-catalyzed hydrolysis (37) and purified by preparative reversed-phase HPLC. The following glucosinolates were analyzed by LC-MS/MS: sinigrin, gluconapin, progoitrin, glucoiberin, glucoraphanin, glucoalyssin, and gluconasturtiin and their related ITCs—AITC, 3-butenyl ITC, 5-vinyloxazolidine-2-thione, 3-methylsulfinylpropyl ITC, sulforaphane, 5-methylsulfinylpentyl ITC, and phenethyl ITC, respectively.
Glucosinolates and ITCs were determined in broccoli seeds by initial heating at 110°C for 2 h (to inactivate myrosinase). The seeds were then ground to a fine powder, lipid was removed by extraction with chloroform, and the residual solid was extracted twice with 75% methanol at 75°C. The combined methanol extracts were concentrated by removal of solvent under reduced pressure, filtered (0.2 μm), spiked with authentic standard analytes, and analyzed by LC-MS/MS. For detection of ITCs in samples of seed extract and in culture medium, ITCs were extracted into dichloromethane and derivatized with ammonia (1.33 mol/L, 24 h at 20°C). The derivatized extracts were then evaporated under reduced pressure, reconstituted in 50% methanol, filtered (0.2 μm), spiked with authentic standard analytes, and analyzed by LC-MS/MS.
Glucosinolates were detected by negative ion electrospray multiple reaction monitoring (MRM), where the fragment ion was hydrogen sulfate (38). Derivatized ITCs were detected by positive ion electrospray MRM, where fragmentation involved loss of ammonia. For LC-MS/MS, the HPLC column was a 100 × 2.1-mm octadecyl silica Symmetry column with a 10 × 2.1-mm guard column (Waters). The flow rate was 0.2 mL/min. The eluent was 0.1% (v:v) trifluoroacetic acid in water, with linear gradients of methanol (0–10% for glucosinolates and 0–80% for ITCs) over 30 min. Source and desolvation temperatures were 120 and 350°C, and the gas flows for cone and desolvation were 150 and 550 L/h, respectively. The capillary voltage was 2.50 kV, and the cone voltage was set at 50 V. Argon gas pressure in the collision cell was 2.9 × 10−3 mbar. Programmed molecular ions, fragment ions, and collision energies were optimized to ±0.1 Da and ±1 eV for MRM detection. Glucosinolate and ITC analytes were quantified by standard addition analysis. Samples analyzed were spiked with 1–100 pmol glucosinolate and 2–100 pmol ITC. The limits of detection for glucosinolates were ≤0.4 pmol and, for ITCs, were ≤2 pmol. The interbatch coefficients of variation were <5%, and recoveries were 80–100%.
Mice feeding experiments
The Ethical Review Committee of the University of Dundee approved this program of work, and, throughout the study, mice were treated as advised by regulations contained in the Animals and Scientific Procedure Act (1986) of the United Kingdom. The nrf2+/+ and nrf2−/− mice were obtained as described previously (33). The mice used in this study have been backcrossed over 6 generations onto a C57BL/6 genetic background. Female mice of between 9 and 14 wk of age were used in all studies. Mice were fed on standard RM1 laboratory feed. Mice were given free access to RM1 feed with broccoli seeds at 15% (by weight) for 7 d immediately before being killed. During the administration of crushed broccoli seed, mice were monitored daily by measurement of body weight. Once the period of feeding these phytochemicals was complete, the mice were killed by exposure to a rising concentration of CO2. Organs were removed and snap-frozen immediately in liquid nitrogen before being stored at −70°C.
Cell culture
Mouse Hepa-1c1c7 cells (European Collection of Animal Cell Cultures) were maintained in minimal essential Eagle's medium, with the Alpha modification (Sigma) supplemented with 10% (v:v) heat-inactivated fetal bovine serum, 50 U/mL penicillin-streptomycin mixture, and l-glutamine at 2 mmol/L. Rat liver RL-34 cells [Japanese Cancer Research Resources Bank (Setagaya-ku)] were grown in Dulbecco's modified Eagle's medium (Life Technologies) supplemented as described above. Wild-type and Nrf2-null mouse embryonic fibroblasts (MEF) were prepared from nrf2+/+ and nrf2−/− mouse lines as described by Tiemann and Deppert (39). These cells were maintained in tissue culture flasks coated with 0.1% (w:v) gelatin for 30 min before use and were grown in medium supplemented with 10 μg/L human recombinant epidermal growth factor, 1 × insulin-transferrin-selenium (Gibco), and 10% (v:v) fetal bovine serum. All cell lines were maintained at 37°C and 5% CO2.
The RL-34, Hepa-1c1c7, and MEF cells were cultured in monolayers and were allowed to grow to 80% confluence in 60-mm dishes before exposure for 24 h to phytochemicals. AITC and sulforaphane were both used to treat cells at a dose of 5 μmol/L. The aqueous broccoli seed extract used to treat cells contained several ITCs, with the total level in the culture media amounting to 0.6 μmol/L.
Enzyme assays and Western blotting
NQO1 enzyme activity was estimated by measuring the dicoumarol-inhibitable fraction of dichlorophenol indophenol reductase activity. GST enzyme activity was measured using 1-chloro-2,4-dinitrobenzene. Western blotting using antibodies against NQO1; class Apha, Mu, and Pi GST isoenzymes; and GCLC subunits was conducted as reported previously (12,19,31).
DNA transfection and luciferase reporter gene assays
Transfection and ARE-reporter gene assays were performed in Hepa-1c1c7 cells. The wild-type mouse nqo1 promoter reporter construct, containing the functional ARE (5′-TCACAGTGAGTCGGCAAAATT-3′) in the pGL3-Basic luciferase reporter vector, was described previously and was designated −1016/nqo5′-luc (29). The mutant NQO1 reporter construct containing 1016 nucleotides of 5′-upstream nqo1 sequence but with the ARE scrambled (i.e., 5′-TTAGAGATACTAGACCACGTC-3′, with mutated bases in italics) is called Mut1 (29). Transfection of −1016/nqo5′-luc and Mut1 into Hepa-1c1c7 cells was performed using Lipofectin Reagent (Life Technologies), and, in all experiments, the pRL-TK Renilla reporter vector (Promega) was used as an internal control. Renilla and firefly luciferase activities were measured using the Dual-Luciferase Reporter Assay System (Promega).
Results
Glucosinolates present in broccoli seeds
The glucosinolate content of broccoli seeds was examined before their ability to induce gene expression in mammalian cells was examined. Prior heating, to inactivate myrosinase, followed by LC-MS/MS analysis revealed that the seeds contained 38.8 mmol glucosinolates per kg. Glucoiberin accounted for 59% of the total glucosinolate recovered, whereas sinigrin and glucoraphanin accounted for 19 and 8% of the glucosinolates, respectively. Significant amounts of progoitrin, gluconapin, and gluconasturtiin were also detected (Table 1). The structures of these phytochemicals are shown in Figure 1.
Glucosinolates (left) and ITCs (right) obtained from broccoli seeds.
Glucosinolates (left) and ITCs (right) obtained from broccoli seeds.
Glucosinolate content of broccoli seeds
| Glucosinolate | Amount | % |
|---|---|---|
| mmol/kg | ||
| Sinigrin | 7.5 ± 1.0 | 19 |
| Gluconapin | 1.3 ± 0.2 | 3 |
| Progoitrin | 2.7 ± 0.4 | 7 |
| Glucoiberin | 23.2 ± 3.6 | 59 |
| Glucoraphanin | 3.2 ± 0.4 | 8 |
| Glucoalyssin | 0.2 ± 0.04 | 1 |
| Gluconasturtiin | 0.9 ± 0.3 | 2 |
| Glucosinolate | Amount | % |
|---|---|---|
| mmol/kg | ||
| Sinigrin | 7.5 ± 1.0 | 19 |
| Gluconapin | 1.3 ± 0.2 | 3 |
| Progoitrin | 2.7 ± 0.4 | 7 |
| Glucoiberin | 23.2 ± 3.6 | 59 |
| Glucoraphanin | 3.2 ± 0.4 | 8 |
| Glucoalyssin | 0.2 ± 0.04 | 1 |
| Gluconasturtiin | 0.9 ± 0.3 | 2 |
Glucosinolate content of broccoli seeds
| Glucosinolate | Amount | % |
|---|---|---|
| mmol/kg | ||
| Sinigrin | 7.5 ± 1.0 | 19 |
| Gluconapin | 1.3 ± 0.2 | 3 |
| Progoitrin | 2.7 ± 0.4 | 7 |
| Glucoiberin | 23.2 ± 3.6 | 59 |
| Glucoraphanin | 3.2 ± 0.4 | 8 |
| Glucoalyssin | 0.2 ± 0.04 | 1 |
| Gluconasturtiin | 0.9 ± 0.3 | 2 |
| Glucosinolate | Amount | % |
|---|---|---|
| mmol/kg | ||
| Sinigrin | 7.5 ± 1.0 | 19 |
| Gluconapin | 1.3 ± 0.2 | 3 |
| Progoitrin | 2.7 ± 0.4 | 7 |
| Glucoiberin | 23.2 ± 3.6 | 59 |
| Glucoraphanin | 3.2 ± 0.4 | 8 |
| Glucoalyssin | 0.2 ± 0.04 | 1 |
| Gluconasturtiin | 0.9 ± 0.3 | 2 |
Broccoli seeds induce NQO1 and GST in nrf2+/+ but not in nrf2−/− mice
Feeding nrf2+/+ mice diets containing 15% (w:w) crushed broccoli seeds resulted in the induction of both NQO1 and GST enzyme activities in the stomach, the small intestine, and the liver, but no increase was observed in the large intestine.
In the wild-type mice, feeding the seeds increased NQO1 activity in the stomach from 155 ± 40 to 248 ± 50 nmol · min–1 · mg–1 protein; in the small intestine, the broccoli seed diet increased NQO1 activity from 106 ± 16 to 183 ± 8 nmol·min–1·mg–1 protein; and, in the liver, this diet increased NQO1 activity from 50 ± 7 to 72 ± 4 nmol ·min–1 · mg–1 protein. The NQO1 enzyme activity in the stomach, the small intestine, and the liver of nrf2−/− mice placed on a control diet was only 50 ± 16, 40 ± 20, and 7 ± 5 nmol · min–1 · mg–1 protein, respectively. The NQO1 enzyme activity did not appear to be increased in stomach, the small intestine, or the liver of nrf2−/− mice fed diet containing broccoli seeds.
In nrf2+/+ mice, feeding the broccoli seed diet for 7 d increased GST activity in the stomach from 1.55 ± 0.10 to 2.53 ± 0.47 μmol · min–1 · mg–1 protein, in the small intestine from 1.61 ± 0.11 to 2.02 ± 0.23 μmol · min–1 · mg–1 protein, and in the liver from 4.91 ± 0.52 to 7.7 ± 0.95 μmol ·min–1 · mg–1 protein. Not only was transferase activity substantially lower in nrf2−/− mice than in the wild-type mice, but also, it was not increased in the mutant mice fed broccoli seeds. In stomach, small intestine, and liver, GST activity in knockout mice on a control diet was 1.26 ± 0.11, 1.18 ± 0.12, and 1.72 ± 0.69 μmol · min–1 · mg–1 protein, respectively.
The levels of NQO1 protein in the tissues of mice fed broccoli seeds was examined by Western blotting to determine whether increases in oxidoreductase activity in stomach, small intestine, and liver reflected an increase in protein. Immunoblotting showed increases of ∼2-fold in the level of NQO1 in all 3 organs from nrf2+/+ mice administered broccoli seeds (Figs. 2and 3). Similar experiments were carried out using antisera against class Alpha, Mu, and Pi GST subunits. These revealed significant increases of class Alpha GSTA3 protein in stomach and small intestine and a modest increase in all organs of class Mu GSTM1. The level of the class Pi GSTP1 subunit did not appear to increase in mice after administration of broccoli seeds.
Nrf2–dependent induction of NQO1, GST, and GCLC proteins in stomach of mice fed broccoli seeds. Western blotting was performed on tissue extracts. Immunoreactive standards were applied to lane 1. Portions (10 μg protein) from stomach (panel a) or small intestine (panel b) of nrf2+/+ and nrf2−/− mice fed on the RM1 control diet and from mice fed on RM1 diet containing 15% (w:w) crushed broccoli seeds were applied to the remaining lanes. Samples of tissue cytosol from 3 mice were applied to the gel.
Nrf2–dependent induction of NQO1, GST, and GCLC proteins in stomach of mice fed broccoli seeds. Western blotting was performed on tissue extracts. Immunoreactive standards were applied to lane 1. Portions (10 μg protein) from stomach (panel a) or small intestine (panel b) of nrf2+/+ and nrf2−/− mice fed on the RM1 control diet and from mice fed on RM1 diet containing 15% (w:w) crushed broccoli seeds were applied to the remaining lanes. Samples of tissue cytosol from 3 mice were applied to the gel.
Induction of hepatic NQO1 and GST by broccoli seeds. Immunoblotting was performed on hepatic cytosol from wild-type and mutant mice as described. Lactate dehydrogenase (LDH) was used as a loading control for the samples.
Induction of hepatic NQO1 and GST by broccoli seeds. Immunoblotting was performed on hepatic cytosol from wild-type and mutant mice as described. Lactate dehydrogenase (LDH) was used as a loading control for the samples.
Western blots showed a 5-fold increase in the level of GCLC in the stomach of nrf2+/+ mice fed broccoli seeds, and a more modest increase was also observed in the small intestine of wild-type mice (Fig. 2). By contrast, no increase was observed in the liver (Fig. 3). A decrease in GCLC protein levels was seen in the stomach, the small intestine, and the liver of nrf2−/− mice compared with the same organs from wild-type mice. Furthermore, the protein was not induced in either stomach or small intestine of mutant mice fed broccoli seeds (Fig. 2).
Glucosinolate breakdown products identified in broccoli seed extracts
The total amount of glucosinolate in the broccoli seed extract was <3.6 μmol/L, whereas the total amount of ITC in the extract was 596 μmol/L. Table 2 shows that 3-methylsulfinylpropyl ITC and sulforaphane account for 61 and 30%, respectively, of the ITCs present in the seed extract. Significant amounts of AITC and 3-butenyl ITC were also obtained.
Isothiocyanates in broccoli seed extracts
| Parent glucosinolate | Isothiocyanate | Amount of isothiocyanate1 | |
|---|---|---|---|
| μmol/L | % | ||
| Sinigrin | Allyl ITC | 25.0 | 4.2 |
| Gluconapin | 3-Butenyl ITC | 23.0 | 3.8 |
| Progoitrin | 5-Vinyloxazolidine-2-thione | nd | — |
| Glucoiberin | 3-Methylsulfinylpropyl ITC | 364.0 | 61.1 |
| Glucoraphanin | Sulforaphane | 177.0 | 30.0 |
| Glucoalyssin | 5-Methylsulfinylpentyl ITC | 5.5 | 0.9 |
| Gluconasturtiin | Phenethyl ITC | 1.2 | 0.2 |
| Parent glucosinolate | Isothiocyanate | Amount of isothiocyanate1 | |
|---|---|---|---|
| μmol/L | % | ||
| Sinigrin | Allyl ITC | 25.0 | 4.2 |
| Gluconapin | 3-Butenyl ITC | 23.0 | 3.8 |
| Progoitrin | 5-Vinyloxazolidine-2-thione | nd | — |
| Glucoiberin | 3-Methylsulfinylpropyl ITC | 364.0 | 61.1 |
| Glucoraphanin | Sulforaphane | 177.0 | 30.0 |
| Glucoalyssin | 5-Methylsulfinylpentyl ITC | 5.5 | 0.9 |
| Gluconasturtiin | Phenethyl ITC | 1.2 | 0.2 |
nd, not determined.
Isothiocyanates in broccoli seed extracts
| Parent glucosinolate | Isothiocyanate | Amount of isothiocyanate1 | |
|---|---|---|---|
| μmol/L | % | ||
| Sinigrin | Allyl ITC | 25.0 | 4.2 |
| Gluconapin | 3-Butenyl ITC | 23.0 | 3.8 |
| Progoitrin | 5-Vinyloxazolidine-2-thione | nd | — |
| Glucoiberin | 3-Methylsulfinylpropyl ITC | 364.0 | 61.1 |
| Glucoraphanin | Sulforaphane | 177.0 | 30.0 |
| Glucoalyssin | 5-Methylsulfinylpentyl ITC | 5.5 | 0.9 |
| Gluconasturtiin | Phenethyl ITC | 1.2 | 0.2 |
| Parent glucosinolate | Isothiocyanate | Amount of isothiocyanate1 | |
|---|---|---|---|
| μmol/L | % | ||
| Sinigrin | Allyl ITC | 25.0 | 4.2 |
| Gluconapin | 3-Butenyl ITC | 23.0 | 3.8 |
| Progoitrin | 5-Vinyloxazolidine-2-thione | nd | — |
| Glucoiberin | 3-Methylsulfinylpropyl ITC | 364.0 | 61.1 |
| Glucoraphanin | Sulforaphane | 177.0 | 30.0 |
| Glucoalyssin | 5-Methylsulfinylpentyl ITC | 5.5 | 0.9 |
| Gluconasturtiin | Phenethyl ITC | 1.2 | 0.2 |
nd, not determined.
Induction of NQO1 and GST in rodent liver cell lines by broccoli-derived chemicals
Aqueous broccoli seed extracts were used to treat cells at an estimated concentration of total ITC of 0.6 μmol/L in the media. The transformed mouse Hepa-1c1c7 liver cell line and the nontransformed rat liver RL-34 epithelial cells were used in these experiments. The broccoli seed extract increased NQO1 enzyme activity ∼3-fold and ∼5-fold in the Hepa-1c1c7 and RL-34 cells, respectively. Treatment with AITC at 5 μmol/L induced NQO1 catalytic activity ∼2-fold in both Hepa-1c1c7 and RL-34 cells. Treatment with sulforaphane at 5 μmol/L induced NQO1 catalytic activity 4.5-fold and 5.2-fold in Hepa-1c1c7 and RL-34 cells, respectively. By contrast, GST activity was not increased to the same extent in either cell line.
Western blotting showed that the level of NQO1 protein in Hepa-1c1c7 and RL-34 cells (Fig. 4A and B, respectively) grown in normal cell culture medium without the addition of phytochemicals was barely detectable. Treatment of both cell lines with the broccoli seed extract containing a mixture of ITCs substantially increased NQO1 protein. This increase was comparable to the induction of NQO1 protein affected by sulforaphane at 5 μmol/L.
Induction of NQO1, GST, and GCLC by broccoli seed extracts in rodent liver cell lines. Cells were grown in either media alone or for 24 h in media containing sulforaphane (Sul; 5 μmol/L), AITC (5 μmol/L), or 1/1000 dilution of broccoli seed extract. Protein standards were applied to lane 1. The other samples are duplicates of individual treatments taken from separate flasks. Panel A shows data from the transformed Hepa-1c1c7 cells and panel B shows data from the nontransformed rat liver RL-34 cells.
Induction of NQO1, GST, and GCLC by broccoli seed extracts in rodent liver cell lines. Cells were grown in either media alone or for 24 h in media containing sulforaphane (Sul; 5 μmol/L), AITC (5 μmol/L), or 1/1000 dilution of broccoli seed extract. Protein standards were applied to lane 1. The other samples are duplicates of individual treatments taken from separate flasks. Panel A shows data from the transformed Hepa-1c1c7 cells and panel B shows data from the nontransformed rat liver RL-34 cells.
Immunoblots were carried out to determine whether the broccoli extracts induced the various GST subunits. The levels of the class Alpha GSTA1/2 and GSTA3 subunits were found to be increased by the broccoli extract, and the degree of induction was similar to that obtained using sulforaphane.
In both Hepa-1c1c7 and RL-34 cells, the broccoli seed extract induced large increases in GCLC protein (Fig. 4).
Broccoli seed extracts stimulate ARE-driven gene expression
To determine whether broccoli seed extracts can activate gene expression controlled through an ARE enhancer, RL-34 cells were transfected with the mouse −1016/nqo5′-luc reporter construct. Treatment of transfected cells with the standard dose of broccoli seed extract produced a 4.6-fold increase in luciferase activity compared with transfected cells treated with vehicle alone (Fig. 5). By contrast, AITC and sulforaphane, each at 5 μmol/L, produced 1.9-fold and 3.3-fold increases, respectively, in luciferase activity. Similar experiments using a reporter construct driven by the mouse nqo1 promoter that contained a mutant ARE (i.e., Mut1) proved to be unresponsive to broccoli seed extract, AITC, and sulforaphane.
Broccoli seed extracts stimulate ARE–driven gene expression. Mouse Hepa-1c1c7 cells were transfected with a luciferase reporter construct driven by the wild-type mouse nqo1 promoter (−1016/nqo5′-luc) or by the same promoter containing a scrambled ARE (Mut1); pRL-TK Renilla reporter vector was used as an internal control. Sixteen hours after transfection, cells were treated for 24 h with 1/1000 dilution of broccoli seed extract, sulforaphane (Sul; 5 μmol/L), AITC (5 μmol/L), or dimethyl sulfoxide (0.1% v:v).
Broccoli seed extracts stimulate ARE–driven gene expression. Mouse Hepa-1c1c7 cells were transfected with a luciferase reporter construct driven by the wild-type mouse nqo1 promoter (−1016/nqo5′-luc) or by the same promoter containing a scrambled ARE (Mut1); pRL-TK Renilla reporter vector was used as an internal control. Sixteen hours after transfection, cells were treated for 24 h with 1/1000 dilution of broccoli seed extract, sulforaphane (Sul; 5 μmol/L), AITC (5 μmol/L), or dimethyl sulfoxide (0.1% v:v).
Induction of GST and NQO1 by broccoli seed extracts is abolished in nrf2−/− mouse embryonic fibroblasts
To explore whether GST subunits and NQO1 can be induced by broccoli in an Nrf2-dependent fashion, wild-type and mutant MEFs were treated with the seed extract. In the nrf2+/+ MEFs, treatment with the standard dose of broccoli seed extract caused a significant increase in GSTM1, GSTA3, and NQO1. This increase was similar to that seen in wild-type MEFs treated with sulforaphane at 5 μmol/L. In the nrf2−/− MEFs, the levels of GSTM1, GSTA3, and NQO1 were lower than in the wild-type cells, and the seed extract failed to induce these proteins (Fig. 6).
Nrf2-dependent induction of NQO1 and GST in mouse embryonic fibroblasts. Wild-type and Nrf2-null MEFs were derived and maintained as described. In the control treatment group the MEFs were grown in medium supplemented with epidermal growth factor, insulin-transferrin-selenium, and 10% fetal bovine serum. Treatment with broccoli seed extract involved growing MEFs in medium for 24 h with 1/1000 dilution of the filtered aqueous seed extract. The sulforaphane (Sul) treated MEFs were grown for 24 h in the presence of ITC (5 μmol/L).
Nrf2-dependent induction of NQO1 and GST in mouse embryonic fibroblasts. Wild-type and Nrf2-null MEFs were derived and maintained as described. In the control treatment group the MEFs were grown in medium supplemented with epidermal growth factor, insulin-transferrin-selenium, and 10% fetal bovine serum. Treatment with broccoli seed extract involved growing MEFs in medium for 24 h with 1/1000 dilution of the filtered aqueous seed extract. The sulforaphane (Sul) treated MEFs were grown for 24 h in the presence of ITC (5 μmol/L).
Discussion
An increasing body of evidence suggests that high intake of cruciferous vegetables can protect against tumorigenesis (2–6). One mechanism proposed to explain this conclusion is that glucosinolates, which are uniquely abundant in these plants, are converted by the actions of myrosinase to thiol-active metabolites that can stimulate cytoprotective responses in cells of the host (40). The major group of phytochemicals that are believed to stimulate such defenses are ITCs, and these compounds can induce the expression of ARE-driven genes. An alternative hypothesis is that ITCs can cause arrest at the G2/M phase of the cell cycle. This is associated with decreases in cyclin B1, Cdc25B, and Cdc25C proteins (41).
Most of the studies into the cellular effects of glucosinolate-derived compounds have used purified phytochemicals such as sulforaphane, benzyl ITC, and phenethyl ITC (2,14–18). How relevant the doses of pure phytochemical used in such gene induction and cell cycle arrest experiments are to the in vivo situation is unclear. This issue is complicated, because the yield of ITCs from different parental glucosinolates varies substantially and can be influenced significantly by the presence of epithiospecifier protein present in certain crucifers (9,42). In the present paper, broccoli seeds were used as the source of plant glucosinolates because we wished to avoid variations in the content of these chemicals that arise from postgermination metabolism. Furthermore, Fahey et al. (43) reported that the ability of broccoli to induce NQO1 in Hepa-1c1c7 cells diminishes with the age of the plant. Therefore, in this study, we used crushed broccoli seeds in the mice feeding experiments and aqueous seed extracts in the cell culture experiments. Analysis of the glucosinolates revealed that the seeds used in this study contained primarily glucoiberin and sinigrin, with lesser amounts of glucoraphanin and progoitrin (Table 1). In the aqueous broccoli seed extracts, LC-MS/MS revealed the presence of large amounts of ITCs, primarily 3-methylsulfinylpropyl ITC, and sulforaphane (Table 2). The low recovery of AITC in the extracts is noteworthy given the large amount of sinigrin in the broccoli seeds.
Enzyme assay and Western blotting showed that addition of broccoli seeds at 15% (w:w) in the RM1 diet induced NQO1 about 2-fold in stomach, small intestine, and liver of wild-type mice. No induction was observed in the nrf2−/− mice. Similar results were observed by treating the nrf2+/+ and nrf2−/− mouse embryonic fibroblasts with broccoli seed extracts. Because the promoter of mouse nqo1 contains a functional ARE that recruits Nrf2 after treatment with sulforaphane (29), it is highly likely that transcriptional activation of mouse nqo1 caused by preparations of broccoli seed is a direct consequence of ITCs stimulating the basic-region leucine zipper protein to transactivate directly the oxidoreductase gene.
Among GSTs, modest increases of GSTM1 protein were observed in the stomach and the small intestine of wild-type but not of nrf2−/− mice after feeding with broccoli seeds. This diet also produced significant increases of the GSTA3 subunit in the stomach and large increases in the small intestine of wild-type mice. However, no such increases were observed in mutant mice. In MEFs from the wild-type and knockout mice, the Nrf2 dependency of induction of GSTM1 and GSTA3 by broccoli was clearly observed. Both the GSTM1 and GSTA3 subunit genes have been reported to contain an ARE (30,44), and it is likely that Nrf2 mediates induction directly through this enhancer. Chromatin immunoprecipitation experiments are required to confirm this hypothesis.
In the stomach and the small intestine of wild-type mice, substantial increases in GCLC were observed after treatment with the broccoli seed preparations. It is likely that Nrf2 mediates the increase in mouse GCLC and requires the existence of a functional ARE in the gene promoter, because this occurs in the human gene (45). However, the presence of a functional ARE in mouse gclc remains to be established.
Cellular models for screening the cancer chemopreventive properties of phytochemicals have frequently used induction of NQO1 enzyme activity in Hepa-1c1c7 cells (46). Our study revealed that besides NQO1 induction, GSTA1/2, GSTA3, and GCLC are also increased significantly in this transformed cell line by broccoli seed extract and by sulforaphane. Importantly, we also found that in nontransformed RL-34 cells, the seed extract and sulforaphane cause large increases in NQO1 and GCLC proteins. Modest increases in GSTA3 were also observed. Nakamura et al. (13) suggested that measurement of GST activity in RL-34 cells provides a useful assay for identifying potential inducing agents. However, our data suggest that induction of NQO1 in these cells may provide the most sensitive assay to identify chemopreventive phytochemicals, because the Western blots in Figure 4 suggest that a 10-fold increase of the protein can be readily achieved.
Significant variations in the amounts and the types of glucosinolates in different broccoli strains appear to exist (7–9). Because this will result in distinct ITCs being generated by myrosinase from different broccoli strains, these differences in glucosinolate content will also influence the level of induction that can be achieved in the host and also possibly the sensitivity to cell-cycle arrest. The significance of variation in the glucosinolate content of cruciferous vegetables in terms of antioxidant and detoxication gene induction and stimulation of cell-cycle arrest and apoptosis warrants further study.
We thank Jed Fahey for valuable constructive criticism of this work during the AICR conference and for pointing out the confounding effects of epithiospecifier protein on glucosinolate degradation.
LITERATURE CITED
,G.
, , B.
& , A.
() Fruit, vegetables, and cancer prevention: a review of the epidemiological evidence
. 18
:–29
., D.T.H.
, , H.
, , R. A.
, , P. A.
& , G.
() A review of mechanisms underlying anticarcinogenicity by brassica vegetables
. 103
:–129
., G.
, , D. T.
, , H.
& , R. A.
() Brassica vegetables and cancer prevention. Epidemiology and mechanisms
. 472
:–168
., H.
, , S.
, , C.
, , E.
, , G.
, , M.
, , S.
& , F.
() Effects of cruciferous vegetables and their constituents on drug metabolising enzymes involved in the bioactivation of DNA-reactive dietary carcinogens
. 480–481
:–297
., P.
& , J. W.
() Phytochemicals from cruciferous plants protect against cancer by modulating carcinogen metabolism
. 131
:–3033
S., C. C.
, , Y.-M.
& , F.-L.
() Isothiocyanates as cancer chemopreventive agents: their biological activities and metabolism in rodents and humans
. 3
:–255
., G. R.
, , R. K.
& , W. J.
() Glucosinolates and their breakdown products in food and food plants
. 18
:–201
., J. W.
, , A. T.
& , P.
() The chemical diversity and distribution of glucosinolates and isothiocyanates among plants
. 56
:–51
., N. V.
, , J. A.
& , E. H.
() Heating decreases epithiospecifier protein activity and increases sulforaphane formation in broccoli
. 65
:–1281
., Y.
, , P.
, , C. G.
& , G. H.
() A major inducer of anticarcinogenic protective enzymes from broccoli: isolation and elucidation of structure
. 89
:–2403
., N.
, , R. K.
, , J. A.
, , G. R.
, , S.R.R.
& , G.
() Dietary glucosinolates as blocking agents against carcinogenesis: glucosinolate breakdown products assessed by induction of quinone reductase activity in murine hepa1c1c7 cells
. 16
:–1194
., V. P.
, , E. M.
, , M. M.
, , S. A.
, , G. J.
, , R.
, , D. J.
, , G. E.
& , J. D.
() Chemoprevention of aflatoxin B1 hepatocarcinogenesis by coumarin, a natural benzopyrone that is a potent inducer of AFB1-aldehyde reductase, the glutathione S-transferase A5 and P1 subunits, and NAD(P)H:quinone oxidoreductase in rat liver
. 60
:–969
., Y.
, , Y.
, , T.
, , H.
, , A.
, , Y.
, , Y.
, , T.
& , K.
() A glutathione S-transferase inducer from papaya: rapid screening, identification and structure-activity relationship of isothiocyanates
. 157
:–200
., C.
, , I. M.
& , J. D.
() Dietary indoles and isothiocyanates that are generated from cruciferous vegetables can both stimulate apoptosis and confer protection against DNA damage in human colon cell lines
. 61
:–6130
., J. D.
, , V. G.
& , G.
() Potent induction of phase 2 enzymes in human prostate cells by sulforaphane
. 10
:–954
., Y.
, , Y.
, , K.
, , H.
, , T.
, , Y.
, , T.
, , F.
& , K.
, et al. () A sulforaphane analogue that potently activates the Nrf2-dependent detoxification pathway
. 277
:–3463
., Z.-Q.
, , C.
, , B.
, , V.
& , A.-N.T.
() Differential responses from seven mammalian cell lines to the treatments of detoxifying enzyme inducers
. 72
:–2253
., J. W.
, , X.
, , P. M.
, , T. W.
, , I.
, , K. K.
, , P.
& , A.
() Sulforaphane inhibits extracellular, intracellular, and antibiotic-resistant strains of Helicobacter pylori and prevents benzo[a]pyrene-induced stomach tumors
. 99
:–7615
., M.
, , K.
, , M.
, , S. A.
, , C. J.
, , L. I.
, , C. R.
, , C.
& , J. D.
() The cap'n'collar basic leucine zipper transcription factor Nrf2 (NF-E2 p45-related factor 2) controls both constitutive and inducible expression of intestinal detoxification and glutathione biosynthetic enzymes
. 61
:–3307
., R. K.
, , K. H.
, , S.
, , T. W.
, , M.
& , S.
() Identification of Nrf2-regulated genes induced by the chemopreventive agent sulforaphane by oligonucleotide microarray
. 62
:–5203
., J. D.
& , L. I.
() Glutathione and glutathione-dependent enzymes represent a co-ordinately regulated defence against oxidative stress
. 31
:–300
., H.
, , T.
, , F.
, , J. D.
& , M.
() Integration and diversity of the regulatory network composed of Maf and CNC families of transcription factors
. 294
:–12
., T.
, , P. J.
, , H. C.
, , C. S.
& , C. B.
() Increased protein stability as a mechanism that enhances Nrf2-mediated transcriptional activation of the antioxidant response element—degradation of Nrf2 by the 26 S proteasome
. 278
:–4541
., K.
, , N.
, , Y.
, , T.
, , T.
& , M.
() Keap1 regulates both cytoplasmic-nuclear shuttling and degradation of Nrf2 in response to electrophiles
. 8
:–391
., M.
, , K.
, , M.
& , J. D.
() Keap1-dependent proteasomal degradation of transcription factor Nrf2 contributes to the negative regulation of antioxidant response element-driven gene expression
. 278
:–21600
., M.
, , N.
, , K.
, , M.
& , J. D.
() Redox-regulated turnover of Nrf2 is determined by at least two separate protein domains, the redox-sensitive Neh2 degron and the redox-insensitive Neh6 degron
. 279
:–31567
., N.
, , A. T.
, , W. D.
, , M. I.
, , A.
, , M.
, , T. W.
& , P.
() Protection against electrophile and oxidant stress by induction of the phase 2 response: fate of cysteines of the Keap1 sensor modified by inducers
. 101
:–2045
., T.
, , P. J.
& , C. B.
() Regulatory mechanisms controlling gene expression mediated by the antioxidant response element
. 43
:–260
., P.
, , M.
, , K.
, , M.
& , J. D.
() Identification of a novel Nrf2-regulated antioxidant response element in the mouse NAD(P)H:quinone oxidoreductase 1 gene; reassessment of the ARE consensus sequence
. 374
:–348
., I. R.
, , Q.
, , K.
, , M.
& , J. D.
() Expression of the aflatoxin B1-8,9-epoxide-metabolizing murine glutathione S-transferase A3 subunit is regulated by the Nrf2 transcription factor through an antioxidant response element
. 64
:–1028
., J. D.
, , S. A.
, , C. J.
, , M.
, , C.
, , G. J.
, , C. R.
& , M.
() The Nrf2 transcription factor contributes both to the basal expression of glutathione S-transferases in mouse liver and to their induction by the chemopreventive synthetic antioxidants, butylated hydroxyanisole and ethoxyquin
. 28
:–41
., P.
& , J. D.
() Contribution of NAD(P)H:quinone oxidoreductase 1 to protection against carcinogenesis, and regulation of its gene by the Nrf2 basic-region leucine zipper and the Arylhydrocarbon receptor basic helix-loop-helix transcription factors
. 555
:–171
., K.
, , T.
, , S.
, , T.
, , K.
, , Y.
, , T.
, , N.
& , K.
, et al. () An Nrf2/small Maf heterodimer mediates the induction of phase II detoxifying enzyme genes through antioxidant response elements
. 236
:–322
., S. A.
, , Q.
, , M.
, , G. K.
, , L. I.
, , C. R.
, , C. J.
, , C. R.
& , G. J.
, et al. () Loss of the Nrf2 transcription factor causes a marked reduction in constitutive and inducible expression of the glutathione S-transferase Gsta1, Gsta2, Gstm1, Gstm2, Gstm3 and Gstm4 genes in the livers of male and female mice
. 365
:–416
., A. T.
, , J. W.
& , P.
() Chemical structures of inducers of nicotinamide quinone oxidoreductase 1 (NQO1)
. 382
:–448
., J.
, , D.
, , P.
& , R.
() Barbarea verna as a source of 2-phenylethyl glucosinolate, precursor of cancer chemopreventive phenylethyl isothiocyanate
. 72
:–764
., C.
, , R.
, , E.
, , E.
, , O.
, , R.
& , S.
() In vitro antiproliferative activity of isothiocyanates and nitriles generated by myrosinase-mediated hydrolysis of glucosinolates from seeds of cruciferous vegetables
. 48
:–3575
., F. A.
, , R. N.
, , B.
& , G.
() Intact glucosinolate analysis in plant extracts by programmed cone voltage electrospray LC/MS: performance and comparison with LC/MS/MS methods
. 306
:–91
., F.
& , W.
() Immortalization of BALB/c mouse embryo fibroblasts alters SV40 large T-antigen interactions with the tumor suppressor p53 and results in a reduced SV40 transformation-efficiency
. 9
:–1915
., A. T.
, , M. A.
, , R. E.
, , R. J.
& , P.
() Potency of Michael reaction acceptors as inducers of enzymes that protect against carcinogenesis depends on their reactivity with sulfhydryl groups
. 98
:–3409
., S. V.
, , A.
, , A. V.
, , K. L.
, , S. K.
, , R.
, , K. D.
, , L.
& , R.
() Sulforaphane-induced G2/M phase cell cycle arrest involves checkpoint kinase 2-mediated phosphorylation of cell division cycle 25C
. 279
:–25822
., R. J.
& , H. L.
() Interactions of thioglucoside glucohydrolase and epithiospecifier protein of cruciferous plants to form 1-cyanoepithioalkanes
. 21
:–1905
., J. W.
, , Y.
& , P.
() Broccoli sprouts: an exceptionally rich source of inducers of enzymes that protect against chemical carcinogens
. 94
:–10372
., J. H.
, , T.
& , I.
() Selective expression of glutathione S-transferase genes in the murine gastrointestinal tract in response to dietary organosulfur compounds
. 25
:–367
., A. C.
, , H. R.
& , R. T.
() Regulation of gamma-glutamylcysteine synthetase subunit gene expression by the transcription factor Nrf2
. 274
:–33636
., Y. H.
& , J. M.
() Induction of quinone reductase as a primary screen for natural product anticarcinogens
. 382
:–414
.Abbreviations
- allyl isothiocyanate
- antioxidant response element
- glutamate cysteine ligase catalytic
- glutamate cysteine ligase modifier
- glutathione S-transferase
- isothiocyanate
- liquid chromatography with triple quadrupole mass spectrometric detection
- mouse embryonic fibroblast
- multiple reaction monitoring
- NAD(P)H:quinone oxidoreductase 1
- nuclear factor-erythroid 2 p45-related factor 2
Footnotes
Published in a supplement to The Journal of Nutrition. Presented as part of the International Research Conference on Food, Nutrition, and Cancer held in Washington, DC, July 15–16, 2004. This conference was organized by the American Institute for Cancer Research and the World Cancer Research Fund International and sponsored by BASF Aktiengesellschaft; Campbell Soup Company; The Cranberry Institute; Danisco USA Inc.; DSM Nutritional Products, Inc.; Hill's Pet Nutrition, Inc.; Kellogg Company; National Fisheries Institute; The Solae Company; and United Soybean Board. An educational grant was provided by The Mushroom Council. Guest editors for this symposium were Helen A. Norman, Vay Liang W. Go, and Ritva R. Butrum.
Supported by grant 2000/11 from the World Cancer Research Fund and by grant G0000268 from the Medical Research Council of the UK (L.G.H.).
© 2004 The American Society for Nutritional Sciences
Arch Biochem Biophys. Author manuscript; available in PMC 2016 Jul 11.
Published in final edited form as:
Published online 2008 Mar 10. doi: 10.1016/j.abb.2008.02.043
NIHMSID: NIHMS53873
The publisher's final edited version of this article is available at Arch Biochem Biophys
See other articles in PMC that cite the published article.
Abstract
Glutathione S-transferase pi has been shown to reactivate 1-cysteine peroxiredoxin (1-Cys Prx) by formation of a complex. A model of the complex was proposed based on the crystal structures of the two enzymes. We have now characterized the complex of GST pi/1-Cys Prx by determining the Mw of the complex, by measuring the catalytic activity of the GST pi monomer, and by identifying the interaction sites between GST pi and 1-Cys Prx. The Mw of the purified GST pi/1-Cys Prx complex is 50,200 at pH 8.0 in the presence of 2.5 mM glutathione, as measured by light scattering, providing direct evidence that the active complex is a heterodimer composed of equimolar amounts of the two proteins. In the presence of 4 M KBr, GST pi is dissociated to monomer and retains catalytic activity, but the Km value for GSH is increased substantially. To identify the peptides of GST pi that interact with 1-Cys Prx, GST pi was digested with V8 protease and the peptides were purified. The binding by 1-Cys Prx of each of four pure GST pi peptides (residues 41–85, 115–124, 131–163, and 164–197) was investigated by protein fluorescence titration. An apparent stoichiometry of 1 mol/subunit 1-Cys Prx was measured for each peptide and the formation of the heterodimer is decreased when these peptides are included in the incubation mixture. These results support our proposed model of the heterodimer.
Keywords: Glutathione S-transferase pi, 1-Cys peroxiredoxin, Heterodimer
Glutathione S-transferases (GSTs1), which catalyze the nucleophilic attack by the thiol of glutathione on electrophilic substrates, constitute a family of enzymes important in the detoxification of xenobiotics, endogenous compounds, and the products of oxidative stress [,]. The pi isozyme (GST pi), crystallized as a homodimer with a subunit molecular weight of 23,500, is of particular interest because it exhibits diverse roles in mammalian cells: it provides a defense against carcinogenesis, since it catalyzes the inactivation of known carcinogens []; it contributes significantly to the development of resistance to cancer chemotherapy, since GST pi levels increase in tumors and the enzyme metabolizes key anticancer drugs [–]; and it promotes the cellular response to oxidative stress, since GST pi has recently been reported to activate the anti-oxidant enzyme 1-Cys peroxiredoxin [,].
1-Cys peroxiredoxin (1-Cys Prx, Prdx 6, Prx VI, and AOP2), a homodimer with a subunit molecular weight of 25,000, is an anti-oxidant enzyme that catalyzes the reduction of hydroperoxides to alcohols using a strictly conserved cysteine []. The oxidized 1-Cys Prx intermediate must react with another thiol compound to regenerate the sulfhydryl cysteine of active 1-Cys Prx. However, that thiol had not been identified.
Recently, we demonstrated that reactivation of human 1-Cys Prx occurs by formation of a complex between 1-Cys Prx and human GST pi in the presence of glutathione (GSH), in which the two proteins are present in equimolar mixtures []. The complex is dissociated when GSH is removed by dialysis []. Complex production in the presence of GSH is followed by glutathionylation of 1-Cys Prx, and subsequently by the slow formation of an intermolecular disulfide between 1-Cys Prx and GST pi []. The disulfide is then reduced by GSH, regenerating an active 1-Cys Prx [,]. A plausible energy-minimized model of the complex was proposed; this ‘in silico’ model was constructed by docking one subunit from the crystal structure of GST pi with one subunit from the crystal structure of 1-Cys Prx using the program ZDOCKpro 1.0. In the present paper, we evaluate the validity of our proposed model by measuring the molecular weight of the complex, by determining whether GST pi retains catalytic activity as a monomer, and by identifying the contact sites between GST pi and 1-Cys Prx in the active complex. A preliminary version of this study has been presented [11].
Materials and methods
Materials
Reduced GSH, 1-chloro-2,4-dinitrobenzene (CDNB), S-hexylglutathione Sepharose, S-hexylglutathione, imidazole, and chemicals for the preparation of buffers were obtained from Sigma Chemical Co. Nickel–nitrilotriacetic acid agarose (Ni–NTA) was purchased from Qiagen, Inc. 1,6-Hexanediol was supplied by ACROS Organics. Synthetic GST pi peptide fragments were obtained from EZBiolab. Centriplus YM-10 concentrators were from Millipore. Staphylococcal Protease V8 was purchased from Worthington Biochemical Corporation. All chemicals were of reagent grade.
Expression and purification of GST pi and 1-Cys Prx
For the expression of human GST pi, the WT plasmid was transformed into Escherichia coli JM105 and the cells were grown and induced for expression of GST pi []. For the expression of human 1-Cys Prx (Prdx 6), the WT plasmid was transformed in E. coli BL21 (DE3) cells []. Because GST pi does not have a His6-tag, purification of the enzyme was performed using a S-hexylglutathione agarose affinity column, as described previously [11]. In all cases, GST pi was eluted using a buffer containing 2.5 mM S-hexylglutathione. 1-Cys Prx contains a His6-tag at the C-terminus; thus, purification of the protein was performed using a Ni–NTA column, as described previously []. All of the enzymes were purified to homogeneity, yielding a single peptide by N-terminal sequencing on an Applied Biosystems gas-phase sequencer (Model Procise) equipped with an on-line Microgradient Delivery System (Model 140 C) and a computer (Model 610 Macintosh). Each of the enzymes exhibited a single band by SDS–PAGE. All purified GSTs were stored in aliquots at −80 °C, and all purified 1-Cys Prx proteins were stored in aliquots at 4 °C, both enzymes in 0.05 M Tris–Cl buffer at pH 8.0.
Determination of the molecular mass of the enzymes
Light scattering measurements with a miniDAWN laser photometer (Wyatt Technology Corp., Santa Barbara, CA) was used to determine the molecular mass of GST pi, 1-Cys Prx, and the complex. All samples (0.1–0.3 mg/mL) were in 0.05 M Tris–Cl, pH 8.0 and filtered through a 0.02-μm filter before being used and, when indicated, a saturating concentration of GSH was included. The addition of GSH to the enzyme samples was used to determine if this substrate would promote dimerization. Data were collected at a laser wavelength of 690 nm and analyzed using ASTRA software for Windows. The A280 nm was used to determine the protein concentration (E0.1%280 nm = 1.24 for GST pi [], 1.07 for 1-Cys Prx, and 1.16 for the complex []).
Light scattering was also conducted for GST pi samples (0.1–0.4 mg/mL) in 0.05 M sodium–MES buffer (pH 6.5) containing various concentrations of KBr (0–4 M). The average molecular mass of each sample was also determined in the presence of a saturating GSH concentration.
Standard assay for GST activity
Enzymatic activity toward 1-chloro-2,4-dinitrobenzene (CDNB) was measured in a total volume of 1.0 mL using a Hewlett–Packard 8453 spectrophotometer by monitoring the formation of the conjugate of CDNB (3 mM) and GSH (2.5 mM) at 340 nm (Δε = 9600 M−1 cm−1) in 0.1 M potassium phosphate buffer (pH 6.5), containing 1 mM EDTA and 2.5% ethanol at 25 °C, according to the method of Habig et al. []. An enzyme unit is defined as the amount of enzyme that converts 1 μmol/min of CDNB to the conjugate. All measurements were corrected for the spontaneous nonenzymatic rate of formation of the conjugate of GSH and CDNB.
The apparent Km value of GST pi for glutathione was determined at 25 °C by varying the glutathione concentrations (0.01–2 mM) while keeping a constant CDNB concentration (3 mM). Similarly, the apparent Km value for CDNB was determined from a range of concentrations of CDNB (0.01–4.0 mM) at a constant glutathione concentration (2.5 mM) in 0.1 M potassium phosphate buffer containing 1 mM EDTA and 2.5% ethanol (pH 6.5).
The kinetic parameters were also determined for GST pi in 0.05 M sodium–MES (pH 6.5) containing the various concentrations of KBr. For these measurements, the enzyme was stored in 0.05 M sodium–MES (pH 6.5) and was diluted into the assay buffer of 0.05 M sodium–MES (pH 6.5) and KBr immediately before the activity measurements. In determining the Km for CDNB in the presence of KBr, the constant [glutathione] was maintained at 25 mM.
Formation of GST pi/1-Cys Prx complex under standard conditions
Heterodimers were generated in which one subunit was a non-His-tagged GST pi and the other subunit was a His-tagged 1-Cys Prx. The enzymes were incubated together in 20% 1,6-hexanediol in 0.05 M Tris–Cl at pH 8.0 for 2 h at 25 °C to facilitate complete dissociation to monomers [,]. A total of 1 mg of each enzyme in 1 mL of buffer was used. This mixture was then dialyzed overnight (molecular weight cutoff = 10 kDa) against 50 mM Tris–Cl buffer containing 2.5 mM reduced GSH at pH 8.0 and 4 °C to remove 1,6-hexanediol and allow the reformation of dimers. The mixture of two proteins was then loaded onto a Ni–NTA column (1.5 mL) equilibrated in 0.05 M Tris–Cl buffer at pH 8.0 containing 2.5 mM reduced GSH at 4 °C. The enzymes separate because 1-Cys Prx has a His-tag on each subunit and binds tightly to the Ni–NTA column, while the GST pi has no His-tag and does not bind to the column. The heterodimer (in which only one of the two subunits has a His-tag) binds less tightly to the Ni–NTA column than does 1-Cys Prx alone (in which every subunit contains a His-tag). The column was washed with 0.05 M Tris–Cl buffer at pH 8.0, containing 2.5 mM GSH, to remove unbound GST pi. The heterodimer and 1-Cys Prx were separated using a linear gradient from 0.05 M Tris–Cl at pH 8.0 containing 2.5 mM reduced GSH to the same buffer with the addition of 0.3 M imidazole (20 mL of each buffer). (If the column is eluted, with buffer lacking GSH, the heterodimer complex dissociates.) The column eluate was monitored at A280 nm. Fractions of 1 mL were collected, and the GST activity was determined under standard conditions. The three peaks were pooled separately and concentrated to approximately 1 mL using the Centriplus YM-10 centrifugal filter device (molecular weight cutoff = 10 kDa). To remove the imidazole, the heterodimer and 1-Cys Prx were dialyzed against 0.05 M Tris–Cl buffer at pH 8.0 containing 2.5 mM reduced GSH. The activity of the recovered GST pi, as well as the purified heterodimer, was determined under standard conditions. The A280 nm was used to determine the protein concentration. All three enzymes were stored at 4 °C in 0.05 M Tris–Cl buffer at pH 8.0 containing 2.5 mM reduced GSH. The N-terminal sequences of the recovered proteins were determined to confirm the composition and purity of the enzymes.
Proteolytic digestion of GST pi by Staphylococcal protease V8
GST pi (1.0–2.0 mg/mL) was incubated for 30 min in 0.05 M Tris–Cl buffer (pH 8.0) with 6 M Urea and 10 mM N-ethylmaleimide at 25 °C to block the free −SH groups. The enzyme solution was then dialyzed overnight (molecular weight cutoff = 10 kDa) against 4 L of 50 mM ammonium bicarbonate (pH 7.8). The enzyme solution was lyophilized and then resolubilized in 1.0 mL of 2 M urea in 50 mM ammonium bicarbonate (pH 7.8) and incubated at 37 °C for 2 h. V8 protease [5.0% (w/w)] and the enzyme sample were incubated for 2.5 h at 37 °C. A second aliquot of the V8 protease solution was added, and incubation was continued for another 2.5 h at 37 °C.
HPLC separation of peptides
The proteolytic digest was injected onto a Varian 5000 LC HPLC (Varian, Walnut Creek, CA) equipped with a Phenomenex C18 reverse-phase column (0.46 × 25 cm) equilibrated with solvent A (0.1% trifluoroacetic acid in water). After elution with solvent A (0% solvent B) for 10 min, a linear gradient was run to 10% solvent B (0.075% trifluoroacetic acid in acetonitrile) at 30 min followed by a linear gradient to 25% solvent B at 105 min. The linear gradient was continued to 35% solvent B at 115 min, followed by 45% solvent B at 180 min, and finally a 30-min gradient to 100% solvent B. The column was eluted for 10 min with 100% solvent B. The flow rate was 1 mL/min. The eluate was monitored at 220 nm with 1 mL fractions collected. The peptide peaks resulting from the HPLC separation were lyophilized and analyzed by N-terminal sequencing as described previously.
Formation of GST pi/1-Cys Prx complex in the presence of GST pi peptide fragments
GST pi/1-Cys Prx complexes were formed under standard conditions (2 h of incubation with 20% 1,6-hexanediol at 25 °C) as described above except that either a fraction from the separated GST pi proteolytic digest or 100 μM of a pure synthetic GST-derived peptide was included. The GST pi, heterodimer complex and 1-Cys Prx were separated using Ni–NTA chromatography. The conditions were the same as described above, except that dialysis was conducted using a membrane with a molecular weight cutoff of 500 to remove 1,6-hexanediol while retaining the proteins and the test peptides. The activity of the recovered GST pi homodimer, as well as the purified complex, was determined under standard conditions as described above. The A280 nm was used to determine the protein concentration. The N-terminal sequences of the recovered GST pi, 1-Cys Prx, and purified heterodimers were determined to confirm the purity of the enzymes.
Fluorescence spectrophotometry
The steady-state fluorescence spectra of 1-Cys Prx (2 μM enzyme subunits) in 0.05 M Tris–Cl buffer, pH 8.0 were measured on a Perkin-Elmer MPF-3 fluorescence spectrophotometer at 25 °C. The enzyme solutions were excited at 280 nm and the emission spectrum of each sample was scanned and recorded in the range of 290–420 nm. The bandwidth for excitation and emission was 10 nm. The spectra were corrected for the background contributed by the buffer.
The stoichiometry of binding for each synthetic GST peptide fragment was determined by fluorescence titration. The decrease in protein fluorescence of 1-Cys Prx (2 μM) in 0.05 M Tris–Cl buffer, pH 8.0, was monitored as a function of peptide concentration (0.04–4 μM). For all measurements, the excitation wavelength was 280 nm and the emission was monitored at 330 nm at 25 °C. In a typical experiment, two parallel samples containing a 1 mL solution of 1-Cys Prx or buffer alone were titrated with 2 μL aliquots of a concentrated GST pi peptide solution. The final dilution did not exceed 2–3% of the initial volume. Fluorescence titration data were corrected for dilutions.
Protein–protein docking of the GST pi and 1-Cys Prx complex
We have used a two stage procedure called ZDOCKpro 1.0, as described previously []. The ZDOCKpro package is based on the ZDOCK and RDOCK programs within Insight II molecular modeling software (Molecular Simulations, Inc.). The Protein Data Bank structures used were PDB: 1PRX (1-Cys Prx) and 19GS (GST pi). The model of the heterodimer complex is the same one presented previously [].
Results
The complex between GST pi and 1-Cys Prx is a heterodimer
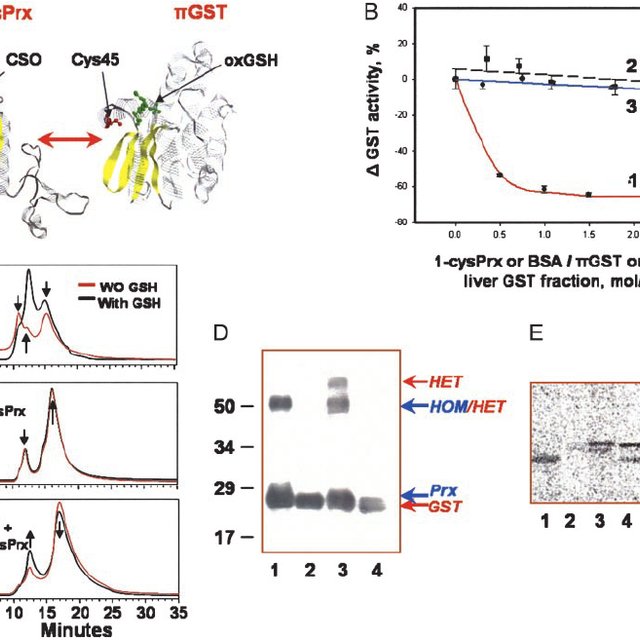
Previously, we isolated and characterized a complex between GST pi and 1-Cys Prx that had a molar ratio of 1:1 based on N-terminal sequencing, but the molecular mass of this complex had not been determined []. In order to assess the composition of the complex, molecular masses for wild-type human GST pi, 1-Cys Prx, and the complex were determined using light scattering (Table 1). First, the molecular mass of wild-type human GST pi was examined over a range of protein concentrations (0.1–0.4 mg/mL) in the absence and presence of glutathione. The monomeric mass of GST pi is 23,500 Da; thus, if the enzyme were completely dimeric in solution, the molecular mass should be approximately 47,000 Da. In contrast, the results shown in Table 1 (columns 2 and 3) indicate that the molecular mass of GST pi is significantly lower than that expected for a fully dimeric species, suggesting that in solution, GST pi exists in a dimer–monomer equilibrium with the monomer predominating.2 The percentage of dimer and monomer in the protein solutions can be determined from the measured average mass at a known protein concentration; the fraction of the total concentration that is monomer and dimer can be calculated using the known mass values of the monomer and the dimer. For example, at a concentration of 0.1 mg/mL of GST pi, the average molecular mass is 29,300 Da in the presence of GSH as compared to 47,000 Da if the protein was 100% dimer, suggesting that the enzyme is only 25% dimeric in solution. The existence of appreciable amounts of GST pi monomer in solution may account, in part, for the ability of this isozyme of GST to form a complex with 1-Cys Prx. For comparison, under similar conditions, the Mw is 50,100 Da for the alpha-GST [] and 46,900 Da for the mu-GST [], indicating that these enzymes are, respectively, 88% and 83% dimeric in solution. These results may explain, in part, the observation that of the glutathione S-transferases tested for the ability to form a complex with 1-Cys Prx, alpha class GST had no effect and mu class was only 20% as effective as GST pi [].
Table 1
Molecular mass of human GST pi, 1-Cys Prx, and the complex in 0.05 M Tris-chloride buffer (pH 8.0), determined by light scattering in the absence and presence of GSH
| [Protein] (mg/mL) | Mass of GST pi (Da) | Mass of 1-Cys Prx (Da) | Mass of GST pi/1-Cys Prx complex (Da) | ||
|---|---|---|---|---|---|
| –GSH | +GSHa | –GSH | +GSHb | +GSHa | |
| 0.1 | 33,200 ± 1800 | 29,300 ± 2100 | 37,400 ± 1000 | 38,700 ± 1100 | 49,300 ± 300 |
| 0.2 | 32,900 ± 1000 | 34,100 ± 2000 | 40,500 ± 400 | 36,900 ± 700 | 49,400 ± 900 |
| 0.3 | 34,300 ± 1400 | 36,300 ± 1800 | 38,800 ± 100 | 37,600 ± 400 | 51,800 ± 4700 |
| 0.4 | 35,100 ± 1300 | 38,700 ± 1800 | 43,000 ± 200 | 35,200 ± 200 | – |
bThe concentration of GSH is 2.5 mM.
Since GSH has been shown to be an important component in the formation and activation of the complex between GST pi and 1-Cys Prx [], the molecular mass of GST pi was also determined in the presence of the same concentration of GSH used to form the complex (2.5 mM). Table 1 (columns 2 and 3) shows that the addition of GSH does not appreciably change the molecular mass of GST pi indicating that GSH does not promote complex formation by shifting the dimer–monomer equilibrium of GST pi.
It was important to ascertain the molecular mass of 1-Cys Prx alone. The 1-Cys Prx subunit (with 6 His) has a molecular mass of 25,900; in this case, if 1-Cys Prx is completely dimeric in solution, the molecular mass should be approximately 52,000. The results of Table 1 (columns 4 and 5) suggest that 1-Cys Prx also exists in an equilibrium mixture of dimer and monomer, as shown by an average molecular mass of about 37,000 Da, significantly lower than the value expected for a dimer. Furthermore, the presence of GSH did not have an effect on the equilibrium. For 1-Cys Prx, the average molecular mass suggests that in solution this protein is 53% dimeric, which also supports the proposal that the observed equilibrium may be partly responsible for its ability to form a complex with another protein.
The molecular weight of the purified GST pi/1-Cys Prx complex was determined only in the presence of GSH because of the instability of the complex in the absence of GSH []. Under these conditions, the average molecular mass of the complex is equal to that of a dimer (~50 kDa; Table 1); the complex cannot be explained by a mixture of 1-Cys Prx + GST pi since the molecular mass is greater than that for either single protein when measured alone. For the first time, we have direct evidence that the active GST pi/1-Cys Prx complex is indeed a dimer.
GST pi is active as a monomer
In our previous studies [], we showed that the complex is active with respect to both peroxidase and glutathione S-transferase activity. However, the GST activity in the complex (when considered per mg of GST present) was only 25% of that of GST pi alone. Since we now know that the complex exists as a heterodimer in which one subunit of GST pi interacts with one subunit of 1-Cys Prx, it is likely that GST pi retains some catalytic activity as a monomer. Furthermore, the glutathione S-transferase activity should properly be compared with that of a monomer of GST. Until now the question of whether the GST pi monomer has activity has not been addressed.
Monovalent anions have been used to disrupt the electrostatic interactions between subunits of multimeric proteins []. KBr was selected for this study since, in accordance with the Hofmeister series of monovalent anions, bromide is an effective protein destabilizing and dissociating reagent []. Moreover, Hearne and Colman [] established that 0.05 M MES (pH 6.5) containing a range of KBr concentrations was a satisfactory system for examining the dimer–monomer equilibrium of the mu-class GST. Therefore, 0.5–4.0 M KBr was added to GST pi in 0.05 M MES (pH 6.5). Table 2 (column 2) shows that the average molecular mass of GST pi in 0.05 M MES (pH 6.5) is dependent on the concentration of KBr present. Appreciable amounts of dimer remain at concentrations up to 2.0 M KBr (Table 2). Above 2.0 M KBr, the average molecular mass decreases as the concentration of KBr increases. The enzyme becomes predominantly monomeric (~94%) in the presence of 4 M KBr (Table 2). The Mw was evaluated in the absence and presence of saturating GSH concentrations to determine whether this substrate promotes dimerization. The results (Table 2, column 3) show that GSH has little effect on the average molecular mass of the enzyme in the presence of KBr.
Table 2
Molecular mass and specific activities of GST pi in 0.05 M MES (pH 6.5) containing various concentrations of KBr, determined by light scattering in the absence and presence of GSH
| Concentration of KBra (M) | Mw without GSH (Da) | Mw with GSHb (Da) | Specific activityb,c (μmol/min/mg) |
|---|---|---|---|
| 0 | 32,400 ± 1000 | 33,800 ± 1000 | 71.6 ± 4.0 |
| 0.5 | 31,500 ± 400 | 32,500 ± 200 | 70.2 ± 4.1 |
| 1.0 | 31,000 ± 1400 | 33,900 ± 700 | 48.0 ± 5.5 |
| 1.5 | 30,700 ± 1900 | 31,400 ± 300 | 32.3 ± 1.0 |
| 2.0 | 30,600 ± 1700 | 30,600 ± 700 | 17.7 ± 1.6 |
| 2.5 | 28,300 ± 1400 | 31,600 ± 1300 | 17.6 ± 2.2 |
| 3.0 | 27,500 ± 900 | 26,100 ± 500 | 13.4 ± 1.3 |
| 3.5 | 25,500 ± 600 | 25,700 ± 2500 | 19.4 ± 1.0 |
| 4.0 | 24,800 ± 2800 | 24,300 ± 2000 | 12.8 ± 1.3 |
aIn all cases, the enzyme (0.1-0.4 mg/mL) was in 0.05 M MES (pH 6.5) containing various concentrations of KBr.
cGST activity, in the presence of various concentrations of KBr, was measured using CDNB as the electrophilic substrate.
GST pi was also assayed for activity in the presence of various concentrations of KBr using CDNB as the electrophilic substrate. As shown in Table 2 (column 4), the specific activity between 0 and 0.5 M KBr is essentially the same, but sharply decreases between 0.5 and 1.0 M KBr. At concentrations greater than 1.0 M KBr the specific activity gradually decreases until the values plateau at 2.0 M KBr (~15 lmol/min/mg). In contrast, most of the decrease in molecular mass of GST pi occurs between 2.0 and 4.0 M KBr (Table 2).
Incubation of GST pi over 5 h in 0.05 M sodium–MES (pH 6.5) containing 4 M KBr indicates that the enzyme, when assayed under the same salt conditions as those in the incubation solution, exhibits the same activity over the entire time period. Since there is no observable time dependence of change in enzymatic activity, we conclude that the activity of the wild-type enzyme in 0.05 M MES (pH 6.5) containing KBr is established rapidly and does not change over at least 5 h.
The kinetic parameters of GST pi were determined in the presence of increasing concentrations of KBr to evaluate whether the electrophilic substrate, the glutathione substrate, or both are affected by the KBr and the dissociation of the dimer. Table 3 shows the Km value for GSH and the Km and Vmax values for the xenobiotic substrate CDNB. For CDNB, the Km values do not change appreciably. In contrast, the Km for GSH exhibits a ~28-fold increase. Some of this Km effect is likely due to the dissociation of GST pi and some due to the effect of KBr on the catalytic reaction, because there is little change in molecular weight up to 2.0 M KBr while the Km value for GSH increases 16-fold. Similar to the specific activity values, the Vmax decreases to approximately 13 μmol/min/mg, clearly indicating that the monomer of GST pi retains appreciable activity.
Table 3
CDNB and GSH kinetic parameters of GST pi in 0.05 M MES (pH 6.5) containing various concentrations of KBr
| Concentration of KBr (M) | aVmaxCDNB (μmol/min/mg) | bKmCDNB (mM) | cKmGSH (mM) |
|---|---|---|---|
| 0 | 71.6 ± 4.0 | 1.0 ± 0.04 | 0.1 ± 0.04 |
| 0.5 | 70.2 ± 4.1 | 1.0 ± 0.1 | 0.4 ± 0.02 |
| 1.0 | 48.0 ± 5.5 | 0.9 ± 0.1 | 0.6 ± 0.1 |
| 1.5 | 32.3 ± 1.0 | 0.6 ± 0.03 | 1.2 ± 0.2 |
| 2.0 | 17.7 ± 1.6 | 0.8 ± 0.01 | 1.6 ± 0.02 |
| 3.0 | 13.4 ± 1.3 | 0.6 ± 0.04 | 2.2 ± 0.1 |
| 4.0 | 12.8 ± 1.3 | 0.6 ± 0.03 | 2.8 ± 0.3 |
aThe Km values were generally determined at 25 °C under saturating conditions of the invariant substrate and the Vmax values were determined by an extrapolation of the velocity vs. the concentration of CDNB to infinite concentrations using SigmaPlot for data analysis. The data presented are the averages of at least two trials.
bThe concentration of GSH used for the determination of the KmCDNB was 25 mM.
cThe concentration of CDNB used in the determination of the KmGSH was 3 mM.
To determine whether the monomer of GST pi has activity, we assumed that the measured Vmax is the sum of the contributions from the dimer and the monomer:
Vmax = x (fraction of total that is dimer) + y (fraction of total that is monomer), where x is the specific activity of the dimer and y is the specific activity of the monomer. We can calculate the fraction of the total that is dimer and monomer from the measured average molecular mass. We selected molecular mass values for the enzyme in the presence of 2.0 M KBr because this activity reflects the effect of KBr on the activity of the enzyme in the absence of any appreciable change in the dimer–monomer equilibrium. Assuming that the activity for the monomer in 2.0 M KBr is the same as the activity of the monomer in the presence of 4.0 M KBr (where GST pi is 94% monomeric), one has two equations and two unknowns; thus a Vmax value can be calculated for the monomer and the dimer of GST pi of 12.0 and 31.1 μmol/min/mg, respectively. The monomer of GST pi is catalytically competent and contributes 28% of the overall specific activity. We have previously reported that the activity of the GST pi subunit within the GST pi/1-Cys Prx heterodimer is 28% that of GST pi alone. Thus, the GST activity in the complex is comparable to that of a separate GST pi monomer suggesting that the GST subunit functions independently in the heterodimer.
Effect of V8 proteolytic fragments of GST pi on the formation of the complex
The heterodimer complex is formed by incubation of purified GST pi (without a His6-tag) and 1-Cys Prx (with a C-terminal His6-tag) at pH 8.0 in buffer containing 20% 1,6-hexanediol to dissociate the homodimers, followed by dialysis against a buffer containing 2.5 mM glutathione (GSH) but lacking l,6-hexanediol []. The heterodimer can be purified by chromatography on a nickel–nitriloacetic acid-agarose column in the presence of GSH as described in Materials and methods. Fig. 1 (open circles) illustrates the separation of the enzymes. As proved by N-terminal sequencing in our earlier paper [], Peak I consists of GST pi and has no His6-tag; Peak II is composed of GST pi/1-Cys Prx heterodimer and it has only one His6-tag; and Peak III consists of 1-Cys Prx, which has two His6-tags/dimer.
A representative plot comparing the separation of dimeric species of GST pi and 1-Cys Prx for the incubation mixture alone (○) and the incubation mixture in the presence of a complete, unfractionated digest of GST pi (●) using a Ni–NTA column as monitored by absorbance at 280 nm. The column was initially (fractions 1–15) eluted with 2.5 mM glutathione in 50 mM Tris–Cl, pH 8.0 (Buffer A). At fraction 15, a linear gradient was started from Buffer A (20 mL) to 0.3 M imidazole in Buffer A (20 mL). After the gradient, elution was continued with 0.3 M imidazole in Buffer A.
We sought to identify the contact sites between GST pi and 1-Cys Prx in the heterodimer. So that we were not restricted to the model we have suggested of the complex [, Fig. 6], we have subjected GST pi to proteolytic digestion by Staphylococcal protease V8 in ammonium bicarbonate buffer, pH 7.8, and tested the peptide peaks for their ability to inhibit the formation of the heterodimer (i.e., Peak II of Fig. 1). Fig. 2 shows the HPLC separation with the peptide peaks numbered. Peaks 1–8 account for the entire sequence of GST pi. The additional peaks likely represent alternate proteolytic cleavages. The molecular weights of most of the peptides range from 650 to 3500. By using a dialysis membrane with a molecular weight cutoff of 500, we have removed the hexanediol (molecular weight ~110) to allow the two proteins to form homo- and heterodimers, yet retained most of the test peptides. The extent of formation of the heterodimer in the presence of the peptide was measured by chromatography on a Ni–NTA column, as in Fig. 1, solid circles.
HPLC separation of GST pi peptides resulting from digestion with Staphylococcal V8 protease. The digest was fractionated on a C18 column, as described under Materials and methods, and the effluent was monitored at A220 nm. The gradient in acetonitrile is shown by the dotted line.
Model of the GST pi/1-Cys Prx heterodimer illustrating the relative locations of the four GST pi peptide fragments that bind to 1-Cys Prx and that inhibit the formation of the complex. Ribbon representation of 1-Cys Prx (PDB 1PRX) complexed with GST pi (PDB 19GS). The backbone of the subunit B of 1-Cys Prx is cyan with tryptophan 33, 82, and 181 highlighted in yellow, and the backbone of the subunit A of GST pi is pink with peptide 41–85 highlighted in blue, peptide 115–124 highlighted in green, peptide 131–163 highlighted in red, and peptide 164–197 highlighted in orange. Glutathione is shown in brown, cysteine 47 of GST pi is green, and cysteine 47 of 1-Cys Prx is yellow.
Table 4 reports the results of these experiments in terms of the area of the heterodimer peak, as measured by A280 nm and by GST pi activity. From the results, it was observed that the heterodimer complex between 1-Cys Prx and GST pi did not form when a complete, unfractionated digest of GST pi was included in the mixture, (compare B with A in Table 4 and the solid with the open circles in Fig. 1). These results indicated that one or more of the GST pi-derived peptides was competing with GST pi and inhibiting the formation of the heterodimer complex. Furthermore, including the mixture of peptides 86–97 + 113–130 (Table 4, Peak 4), peptide 41–85 (Table 4, Peak 6), peptide 131–163 (Table 4, Peak 7), or the mixture of peptides 86–97 and 164–197 (Table 4, Peak 8) in the incubation mixture of GST pi and 1-Cys Prx substantially decreased the formation of the heterodimer complex. In contrast, peptides 32–40 (Table 4, Peak 1), 98–112 + 198–209 (Table 4, Peak 2), 198–209 alone (Table 4, Peak 3) and 1–30 (Table 4, Peak 5) had little effect on the extent of formation of the heterodimer. We thus identified five GST pi peptide regions that may be involved in the interactions between GST pi and 1-Cys Prx complex: residues 41–85, 86–97, 113–130, 131–163, and 164–197. However, for conclusive results, each of these peptides had to be tested separately with defined concentrations. The peptide of residues 115–124 was substituted for 113–130 since, based on the molecular model, this was the region of the peptide making closest contact with 1-Cys Prx. In addition, peptide fragment 32–40 was added to test whether the failure to observe effects for residues 32–36 and 37–40 was due to the fact that the peptides were too small to be retained throughout the workup procedure. These peptides were synthesized for further examination; the sequences of these peptides are shown in Fig. 3. Each of the synthetic peptides was >90% pure.
Sequences of synthetic peptides of GST pi that were examined for interaction with 1-Cys Prx.
Table 4
Yield of GST pi/1-Cys Prx heterodimer in the presence of peptide fragments derived from various regions of GST pia
| GST pi peptide peaks examined | Peak II (heterodimer) | |
|---|---|---|
| Total Abs280 nm units | Total GST activity unitsb (ΔOD340 nm/min) | |
| A. None | 1.18 | 1.36 |
| B. Complete, unfractionated digest | No peak | No peak |
| 1. Residues 32-36 and 37-40 | 1.05 | 1.17 |
| 2. Residues 98-112 and 198-209 | 0.87 | 0.91 |
| 3. Residues 198-209 | 1.09 | 1.21 |
| 4. Residues 86-97 and 113-130 | 0.21 | 0.33 |
| 5. Residues 1-30 | 1.12 | 1.29 |
| 6. Residues 41-85 and V8 peptide | No peak | No peak |
| 7. Residues 131-163 | 0.37 | 0.46 |
| 8. Residues 86-97 and 164-197 | 0.62 | 0.54 |
aGST pi and 1-Cys Prx (40 μM of each) together with each GST pi peptide peak were treated with 20% 1,6-hexanediol, followed by dialysis against buffer containing 2.5 mM GSH and loaded onto a Ni-NTA column, as described in Materials and methods.
bGST activity was measured using CDNB as the electrophilic substrate, as described in Materials and methods, and expressed as AOD340 nm/min.
Effect of pure, synthetic fragments of GST pi on the formation of the complex
The influence of the synthetic peptides of GST pi on heterodimer formation was investigated by including 100 μM of each peptide in the incubation mixture of GST pi and 1-Cys Prx. The extent of formation of the heterodimer in the presence of the peptide was measured by chromatography on a Ni–NTA column, as in Fig. 1. The results are summarized in Table 5. Peptides 41–85 and 115–124 completely inhibited complex formation (Table 5, lines 2 and 3), peptides 131–163 and 164–197 partially inhibited the formation of the complex (Table 5, lines 4 and 5), and peptides 32–40 and 86–97 did not appreciably affect formation of the complex (Table 5, lines 6 and 7). These results agree with the model of the heterodimer complex [] since the peptides that substantially decreased complex formation are close to the subunit interface and suggest they may bind to 1-Cys Prx.
Table 5
Yield of heterodimer protein peak from the Ni-NTA column in the presence of various synthetic GST pi peptidesa
| GST pi peptides | Peak II (heterodimer) | |
|---|---|---|
| Total Abs280 nm units | Total GST activity unitsb (ΔOD340 nm/min) | |
| 1. None | 1.18 | 1.36 |
| 2. Residues 41-85 | No peak | No peak |
| 3. Residues 115-124 | No peak | No peak |
| 4. Residues 131-163 | 0.49 | 0.62 |
| 5. Residues164-197 | 0.66 | 0.74 |
| 6. Residues 32-40 | 1.10 | 1.21 |
| 7. Residues 86-97 | 0.94 | 1.09 |
aGST pi and 1-Cys Prx proteins (40 μM of each) together with 100 μM of synthetic GST pi peptide were treated with 20% 1,6-hexanediol followed by dialysis against buffer containing 2.5 mM GSH and loaded onto a Ni-NTA column, as described in Materials and methods.
bGST activity was measured using CDNB as the electrophilic substrate, as described in Materials and methods, and expressed as ΔOD340 nm/min.
Effect of GST pi peptide fragments on the fluorescence of 1-Cys Prx
1-Cys Prx contains three tryptophans per monomer. Based on the proposed model of the complex, tryptophan 33, 82, and 181 are near the postulated interface; these tryptophans may be spectroscopically affected upon the binding of the GST pi fragments. When we compared the emission spectrum of 1-Cys Prx (Fig. 4, spectrum a) with that of free tryptophan (Fig. 4, spectrum d), we observed a shift in the maximum emission peak likely due to shielding of the tryptophan residues in the proteins from the aqueous phase [].
The steady-state fluorescence spectra of 1-Cys Prx (2 μM enzyme subunits) in 0.05 Tris–chloride buffer (pH 8.0) was measured on a Perkin-Elmer MPF-3 fluorescence spectrophotometer. Emission spectrum (excitation at 280 nm, bandwidth of 10 nm) of 1-Cys Prx alone (a), 1-Cys Prx in the presence of 3.4 μM GST pi fragment 131–163 (b), 1-Cys Prx in the presence of 3.4 μM GST pi fragment 41–85 (c), free tryptophan (6 μM) (d).
A comparison of the emission spectrum (with excitation at 280 nm) of 2 μM of 1-Cys Prx alone (Fig. 4, spectrum a) with that of the same concentration of 1-Cys Prx in the presence of 3.4 μM GST pi peptide 41–85 (Fig. 4, spectrum c) reveals a decrease in the relative fluorescence intensity of 1-Cys Prx from 45 at its emission maximum (330 nm), to ~14 in the presence of fragment 41–85, a reduction to 31% of its original value of 1-Cys Prx caused by the addition of peptide 41–85. In contrast, when the emission spectrum of 1-Cys Prx is measured with the same concentration of GST pi fragment 131–163 (3.4 μM) (Fig. 4, spectrum b), the fluorescence intensity is reduced to only 29 fluorescence units, or to 65% of the fluorescence intensity of 1-Cys Prx alone. The decrease in the fluorescence intensity at the emission maximum of 1-Cys Prx in the presence of peptide 41–85 is perhaps due to a further change in solvent exposure of the tryptophan residues in the enzyme or to quenching by a proximate amino acid; but with fragment 131–163, a smaller change in tryptophan fluorescence is observed.
The binding of GST pi residues 32–40, 41–85, 86–97, 115–124, 131–163, and 164–197 to 1-Cys Prx in 0.05 M Tris–Cl, pH 8.0 was studied by monitoring the effect of increasing concentrations of each peptide on the fluorescence intensity at 330 nm of 1-Cys Prx (2 μM subunit concentration), as described in Materials and methods. Examples of these titrations are shown in Fig. 5A and B. From the concentration of the peptide at which the ΔF/ΔFmax first reaches 1.0 (see Fig. 5A), a stoichiometry was estimated of about 1 mol of peptide bound per mol of 1-Cys Prx subunit for GST pi residues 41–85 and 115–124 (Table 6, lines 1 and 2); in these cases ΔFmax is ~32. Furthermore, a stoichiometry of about 1.0 mol of peptide per enzyme subunit is obtained for GST pi residues 131–163 (see Fig. 5B) and 164–197 (Table 6, lines 3 and 4). However, in these cases ΔFmax is ~15. Under similar conditions, 1-Cys Prx does not bind peptides 32–40 and 86–97 (Table 6 lines 5 and 6), as demonstrated by their failure to change the fluorescence of 2 μM 1-Cys Prx when added at concentrations as high as 20 μM; these experiments indicate the specificity of the designated GST pi peptides in their interactions with 1-Cys Prx.
Fluorescence determination of the stoichiometry of binding for synthetic GST pi fragments 41–85 (A) and 131–163 to 1-Cys Prx (B). 1-Cys Prx (2 μM subunit concentration) in 0.05 M Tris–chloride buffer (pH 8.0) was titrated with the pure GST peptides at 25 °C. The results of the titration are plotted as fraction of maximum change (ΔF/ΔFmax) vs. the concentration of GST pi peptide 131–163. The decrease in protein fluorescence emission (ΔF) was recorded at 330 nm (excitation at 280 nm). The maximum change in fluorescence units (ΔFmax) is 32 for peptide 41–85 and 18 for peptide 131–163.
Table 6
Determination of the stoichiometry of binding for synthetic GST pi peptide fragments and 1-Cys Prxa
| GST pi peptides | Maximum change in fluorescence units (ΔF) | Bound peptide/enzyme subunit |
|---|---|---|
| 1. Residues 41-85 | 32 | 1.10 |
| 2. Residues 115-124 | 35 | 0.95 |
| 3. Residues 131-163 | 18 | 0.95 |
| 4. Residues 164-197 | 11 | 0.73 |
| 5. Residues 32-40 | 0 | 0 |
| 6. Residues 86-97 | 0 | 0 |
aThe stoichiometry for the peptides was determined from the protein fluorescence titrations, as illustrated by Fig. 5. The titration experiments were conducted in 0.05 M Tris-Cl buffer, pH 8.0 at 25 °C at the concentration of enzyme subunits of 2 μM.
It is unclear whether the peptide binding sites for 1-Cys Prx are mutually exclusive or distinct. To distinguish between these possibilities, the fluorescence intensity at 330 nm of 1-Cys Prx (2 μM subunit concentration) was measured in the presence of 10 μM GST pi peptide 41–85 along with 10 μM of either peptides 115–124, 131–163, or 164–197. The combination of 41–85 with either 115–124 or 164–197 yields an average ΔFmax of ~31, similar to the ΔFmax of peptide 41–85 alone (Table 6, line 1). In contrast, the peptide mixture of 41–85 + 131–163 exhibits a slightly larger ΔFmax ~ 38. Moreover, the addition of 10 μM of all the peptides to 1-Cys Prx also yielded a value for ΔFmax ~ 38. These results suggest that peptide fragment 131–163 may bind at a different location from all the other fragments, consistent with the original model of the complex.
Discussion
In previous studies [], we proposed that the mechanism of 1-Cys peroxiredoxin reactivation initially requires heterodimer formation between GST pi and 1-Cys Prx in the presence of glutathione. The formation of the complex allows GSH to have access to the oxidized cysteine 47 of 1-Cys Prx, facilitating its glutathionylation. The GST-bound GSH is thus functioning like the “resolving cysteine” residue in the 2-Cys peroxiredoxins []. Glutathionylation of 1-Cys Prx changes the conformation of the heterodimer, permitting a disulfide to form slowly between Cys-47 of GST pi and Cys-47 of 1-Cys Prx. (However, the intersubunit disulfide is not required for the formation or stabilization of the heterodimer since a heterodimer complex can be isolated chromographically even when S-methyl glutathione replaces reduced glutathione []; in that case, though, the processes of glutathionylation, inter-subunit disulfide formation and reactivation of 1-Cys peroxiredoxin do not occur). In contrast, in the presence of GSH, the intersubunit disulfide is reduced by GSH with regeneration of the peroxidative cysteine 47 of 1-Cys Prx [].
We have determined that the GST pi/1-Cys Prx complex exists as a dimer. This conclusion comes from the observation that the molecular mass of the complex is ~50,000 Da in the presence of GSH [i.e., the sum of the mass of one subunit of GST pi (23,500) and one subunit of 1-Cys Prx (25,900)]. While GST pi and 1-Cys Prx have both been crystallized as dimers, our experiments show that, in solution, the individual proteins exist as mixtures of dimer and monomer since the average molecular mass of the individual proteins is significantly lower than that predicted for dimers. Indeed, the existence of insolution of appreciable amounts of monomeric GST pi and of monomeric 1-Cys Prx make it possible to form a heterodimic complex.
In our previous paper [], we proposed that hexanediol was needed in the heterodimer incubation mixture to dissociate the enzymes to monomers. In light of the results of the present study, we suggest that hexanediol is actually needed to facilitate conformational changes that allow heterodimer formation. In past experiments involving the use of hexanediol with GST A1-1, the organic solvent was apparently needed mainly to dissociate the dimers.
The observation that the heterodimer complex is strictly dimeric in solution, while GST pi and 1-Cys Prx exist in dimer–monomer mixtures, also suggests that the stability of the complex is greater than that of either the dimer of GST pi or the dimer of 1-Cys Prx. This conclusion is further supported by the fact that Peak II is the largest peak in Fig. 1 after dissociation and reformation of the dimers. Although the reformation of dimers from a mixed pool of subunits is a random event, the greater stability of the heterodimer favors that species (i.e., Kd is lowest for the heterodimer).
There are several observations suggesting that GST pi is catalytically active as a monomer. For example, the active GST pi/1-Cys Prx complex, now identified as a heterodimer, has glutathione S-transferase activity. Additionally, although GST pi has been crystallized as a dimer, the sites for glutathione and for small electrophilic substrates (such as CDNB) are predominantly contained within each subunit; thus it is reasonable that monomers, produced under mild conditions, have activity. In a previous study, a monomeric species of a human GST pi was engineered by introducing 10 site specific mutations []. However, the absence of catalytic activity in this structurally stable monomer may not have general implications since it was so drastically altered.
We have here used 0.05 M MES, pH 6.5 containing a range of KBr concentrations to examine the dimer–monomer equilibrium of GST pi. Based on the crystal structure analyses of GSTs, three major interactions between the subunits in the dimer have been identified [–]. These interactions are the electrostatic and hydrogen-bonds between the polar amino acid residues of one subunit and that of the other subunit, the stacking interaction of two symmetrically equivalent arginines (one from each subunit), and the hydrophobic interactions, including a ball-and-socket motif. Electrostatic interactions of multimeric proteins can be disrupted by the addition of salts [26]. For example, the addition of salt resulted in the modulation of the dimer–monomer equilibrium of beta-lactoglobulin at pH 3 [] and of GST mu at pH 6.5 [] due to a shielding of the attractive charged groups in the proteins. We speculate that the addition of KBr also shields the electrostatic attractive forces at the GST pi subunit interface, thus, shifting the dimer–monomer equilibrium toward monomer. We found that the largest change in the specific activity was observed between 0.5 and 2.0 M KBr, while the largest change in the enzyme's average mass is observed between 2.0 and 4.0 M KBr. Therefore, the decrease in the enzyme's activity is primarily due to an effect of KBr on the catalytic reaction, and not to the dissociation of the subunits. The kinetic parameters of GST pi determined in the presence of increasing concentrations of KBr indicate that the binding site that is significantly affected by the separation of the subunits and the ionic interactions is the glutathione site rather than the electrophilic site. At the glutathione site, there is an extensive network of interactions between GSH and the amino acids of the enzyme, most of which involve residues from the same subunit binding the GSH []. However, there is a specific attractive interaction between the α-amino group of glutathione and the carboxylate of Asp98 from the neighboring subunit []. When the enzyme subunits are separated, Asp98 from the opposite subunit is unavailable and the Km for GSH increases. In contrast, the site for CDNB is defined entirely by the side chains of residues within the same subunit []; thus separation of the subunits does not affect the binding by GST pi of this xenobiotic substrate. Furthermore, as indicated in the results from the effect of KBr on Vmax, the GST pi monomer is catalytically competent; in fact, the Vmax of the dimer is only 2.6 times that of the monomer.
A major aim of this work was to identify segments of GST pi that belong to the interface of the complex. It has long been known that fluorescence spectroscopy can be a sensitive approach for investigating the interaction of proteins with target peptides. We have used this tool to probe the interaction of 1-Cys peroxiredoxin with several peptides which could encompass the interaction sites between GST pi and 1-Cys Prx. Tryptophan 33, 82, and 181, highlighted in the model of the GST pi/1-Cys Prx, are in the vicinity of the interface of the complex (Fig. 6), which enabled us to monitor the interaction between the GST peptides and 1-Cys Prx by means of the intrinsic fluorescence of these chromophores. These tryptophan residues of 1-Cys Prx exist in a solvent-restricted or hydrophobic environment as shown by the blue shift in the maximum emission peak observed with 1-Cys Prx when compared to free tryptophan []. In the presence of four GST pi peptides, a decrease in the maximum fluorescence of 1-Cys Prx at 330 nm is observed as compared to the 1-Cys Prx alone, which suggests that these fragments interact with 1-Cys Prx, further shielding it from the solvent or quenching the tryptophans []. Tyrosines and/or aspartates, for example, which are present in these GST pi fragments (see Fig. 3), can account for the quenching in fluorescence that is observed.
The identification of the GST pi segments that bind to 1-Cys Prx and inhibit the formation of the complex allows for the prediction of the types of interactions that could lead to subunit recognition and stabilization of the heterodimer. Based on the model of the complex (Figs. 6 and and7A)7A) and analysis using the ZDock program, which thoroughly analyzes the most favorable protein–protein interaction sites, hydrophobic forces could be the major interactions between the two proteins. For example, GST pi fragment 115–124 is close to fragment 163–169 of 1-Cys Prx. Tyr118 of GST pi is thus only 3.5 Å away from Val163 of 1-Cys Prx, a distance sufficiently close for a hydrophobic interaction. Moreover, Val119 of GST pi is situated 4.1 Å from Val164 of 1-Cys Prx, reinforcing the likelihood of hydrophobic interactions between the two types of subunit. In the case of GST pi fragment 41–85, the aromatic ring of Tyr49 of GST pi is 3 Å from that of Tyr149 of 1-Cys Prx. This distance is sufficiently close for a pi–pi stacking interaction (less than 5.6 Å) []. Furthermore, the phenolic ring of Tyr63 of GST pi is 4.4 Å from the benzyl ring of Phe43 of 1-Cys Prx, also suggesting these two residues may interact through pi–pi stacking. Electrostatic interactions between the two enzymes may also be important. For example, the negatively charged carboxylate of Asp171 of GST pi is 4.2 Å from the positively charged amino group of Arg219 of 1-Cys Prx. Additionally, the imidazole group of His162 in GST pi is situated 4.9 Å from the carboxyl group of Asp180 of 1-Cys Prx. Therefore, if the pK of the Histidine is high enough, these two residues may interact electrostatically.
Model of the GST pi/1-Cys Prx heterodimer, and crystal structures of homodimeric 1-Cys Prx (PDB 1 PRX) and homodimeric human GST pi (PDB 19GS). (A) Heterodimer. The backbone of the 1-Cys Prx subunit is shown in cyan with peptides which includes regions close (<6 Å) to GST pi highlighted in white (left to right): 43–49, 149–155, 160–170 (helix), 176–184, and 218–221. The backbone of the GST pi subunit is shown in pink, with the peptides tested for interaction with 1-Cys Prx highlighted in blue (41–85), green (115–124), red (131–163), and orange (164–197), as in Fig. 6. Glutathione is shown in brown. (B) 1-Cys Prx homodimer. The backbone of subunit B is cyan with the peptides close to GST pi in the heterodimer highlighted in yellow: 43–49, 149–155, 160–170, 176–184, and 218–221. The backbone of subunit A is gold, with the peptides close to GST pi in the heterodimer pictured in white: 218–221, 176–184, 160–170, 149–155, and 43–49. (C) GST pi homodimer. The backbone of subunit B is yellow with the peptides tested for interaction with 1-Cys Prx highlighted in cyan (41–85), green (115–124), red (131–163), and orange (164–197). The backbone of subunit A is pink, with the peptides tested highlighted in blue (41–85), green (115–124), red (131–163), and orange (164–197).
Fig. 7 shows the crystal structures of homodimeric human 1-Cys Prx and GST pi as compared to the proposed model of the heterodimer (Fig. 7A); in this heterodimer, on the 1-Cys Prx subunit (cyan) we have highlighted in white the peptides considered close to GST pi. Fig. 7B depicts 1-Cys Prx with the two subunits shown in cyan and gold, respectively, and with peptides in close contact with GST pi in the heterodimer highlighted in yellow and white. Interestingly, the amino acids that appear to be part of the heterodimer interface are also part of the subunit interface of the 1-Cys Prx homodimer. In contrast, when we analyze the crystal structure of homodimeric human GST pi (Fig. 7C), with subunits shown in pink and yellow, and highlight the major peptides (shown in the same color as in Fig. 7A) that are in close contact with 1-Cys Prx in the heterodimer, most of the amino acids that appear to be part of heterodimer interface are not part of the subunit interface of the GST pi homodimer. The red, green and orange peptide regions are far from the interface of the GST pi subunits (Fig. 7C). Only a small section of the peptide 41–85 (highlighted in cyan on one subunit and blue on the other subunit) makes close contact in the GST pi homodimer (Fig. 7C). We suggest that GST pi predominantly utilizes a different interface for other protein–protein interactions that is distinguishable from the homodimeric interface. GST pi appears to form complexes with other proteins such as Jun N-terminal kinase [–] and tumor necrosis factor alpha []. If the other proteins interact entirely with the same site of GST pi occupied by 1-Cys Prx, complexes with these proteins seem likely to form with a GST monomer. In the case of the GST pi/1-Cys Prx, the subunit interface of 1-Cys Prx is the same for another subunit of 1-Cys Prx or for a GST pi subunit, thus allowing only one subunit of 1-Cys Prx to interact with one subunit of GST pi.
The involvement of GST pi in protein–protein interactions constitutes an important new role for GST pi and must affect the manner in which GST pi influences the cellular response to oxidative stress. Furthermore, since both enzymes are overexpressed in tumor cells, understanding this important interaction may lead to an innovative approach for the development of peptidomimetic agents that may inhibit the formation of the complex in patients undergoing cancer radiation therapy. Radiation therapy for cancer results in the local formation of peroxides that injure the tumor cells, to the benefit of the patients. On the other hand, active GST pi/1-Cys Prx complex may rapidly decompose the peroxides, thereby interfering with the effects of radiation therapy. In this case, inhibitors of complex formation could be a useful adjuvant to radiation therapy.
In summary, we have examined the interaction of glutathione S-transferase pi with 1-Cys peroxiredoxin in order to increase knowledge of the chemical and structural basis of the activation by GST pi of 1-Cys Prx. Three main characteristics were identified: first, the complex is a heterodimer. This led to the second characteristic: GST pi is active as a monomer. Finally, we identified GST pi residues 41–85 and 115–124 as the critical contact sites between GST pi and 1-Cys Prx.
Acknowledgments
We thank Chati Lum Zony for her contributions to the initial experiments and Dr. Yu Chu Huang for N-terminal sequencing of the peptides and proteins. We also thank Dr. Sheldon Feinstein for supplying the extracts of E. coli harboring recombinant human 1-Cys Prx (Prdx 6). The work was supported by NIH Grant R01-CA66561 (R.F.C.), by NIH fellowship 1F31 GM 75387 (L.A.R.) and by NIH Grant P01-HL 79063 (A.B.F.).
Footnotes
1Abbreviations used: GST, glutathione S-transferase; GST pi, pi-class glutathione S-transferase; 1-Cys Prx, 1-Cysteine peroxiredoxin; Ni–NTA, nickel–nitriloacetic acid agarose; TRIS, tris (hydroxymethyl) aminomethane; EDTA, disodium ethylenediamine tetraacetate; MES, 2-(N-morpholino) ethane-sulfonate; WT, wild-type; GSH, glutathione; CDNB, 1-chloro-2,4-dinitrobenzene; SDS–PAGE, sodium dodecyl sulfate–polyacrylamide gel electrophoresis.
2In a separate study, Y.-C. Huang, S. Misquitta, S.Y. Blond, R.F. Colman (manuscript in preparation) have shown that when the protein concentration range is extended from 0.008 to 1.2 mg/ml, the average mass of GST pi changes from 25,500 to 47,000. Furthermore, the substrate CDNB has little influence on the molecular mass.
References
1. Mannervik B, Danielson UH. CRC Crit. Rev. Biochem. 1988;23:283–337. [PubMed] [Google Scholar]
2. Armstrong RN. Chem. Res. Toxicol. 1997;10:2–18. [PubMed] [Google Scholar]
3. Coles B, Ketterer B. Crit. Rev. Biochem. Mol. Biol. 1990;25:47–70. [PubMed] [Google Scholar]
4. Lee FY, Sciandra J, Siemann DW. Biochem. Pharmacol. 1989;38:3697–3705. [PubMed] [Google Scholar]
5. Moscow JA, Fairchild CR, Madden MJ, Ransom DT, Wieand HS, O'Brien EE, Poplack DG, Cossman J, Myers CE, Cowan KH. Cancer Res. 1989;49:1422–1428. [PubMed] [Google Scholar]
6. Nakagawa K, Saijo N, Tsuchida S, Sakai M, Tsunokawa Y, Yokota J, Muramatsu M, Sato K, Terada M, Tew KD. J. Biol. Chem. 1990;265:4296–4301. [PubMed] [Google Scholar]
7. Tsuchida S, Sato K. Crit. Rev. Biochem. Mol. Biol. 1992;27:337–384. [PubMed] [Google Scholar]
8. Manevich Y, Feinstein SI, Fisher AB. Proc. Natl. Acad. Sci. USA. 2004;101:3780–3785.[PMC free article] [PubMed] [Google Scholar]
9. Ralat LA, Manevich Y, Fisher AB, Colman RF. Biochemistry. 2006;45:360–372. [PubMed] [Google Scholar]
10. Choi HJ, Kang SW, Yang CH, Rhee SG, Ryu SE. Nat. Struct. Biol. 1998;5:400–406. [PubMed] [Google Scholar]
11. Ralat LA, Misquitta SA, Zony CL, Manevich Y, Fisher AB, Colman RF. FASEB J. 2007;21:651–653.[Google Scholar]
12. Ralat LA, Colman RF. J. Biol. Chem. 2004;279:50204–50213. [PubMed] [Google Scholar]
13. Chen JW, Dodia C, Feinstein SI, Jain MK, Fisher AB. J. Biol. Chem. 2000;275:28421–28427. [PubMed] [Google Scholar]
14. Pettigrew NE, Brush EJ, Colman RF. Biochemistry. 2001;40:7549–7558. [PubMed] [Google Scholar]
15. Habig WH, Pabst MJ, Jakoby WB. J. Biol. Chem. 1974;249:7130–7139. [PubMed] [Google Scholar]
16. Misquitta SA, Colman RF. Biochemistry. 2005;44:8608–8619. [PubMed] [Google Scholar]
17. Vargo MA, Nguyen L, Colman RF. Biochemistry. 2004;43:3327–3335. [PubMed] [Google Scholar]
18. Hearne JL, Colman RF. Biochemistry. 2006;45:5974–5984. [PubMed] [Google Scholar]
19. Cacace MG, Landau EM, Ramsden JJ. Q. Rev. Biophys. 1997;30:241–277. [PubMed] [Google Scholar]
20. Matyus L, Szollosi J, Jenei A. J. Photochem. Photobiol. B. 2006;83:223–236. [PubMed] [Google Scholar]
21. Wood ZA, Schroeder E, Harris JR, Poole LB. Trends Biochem. Sci. 2003;28:32–40. [PubMed] [Google Scholar]
22. Abdalla AM, Bruns CM, Trainer JA, Mannervik B, Stenberg G. Protein Eng. 2002;15:827–834. [PubMed] [Google Scholar]
23. Ji X, Zhang P, Armstrong RN, Gilliland GL. Biochemistry. 1992;31:10169–10184. [PubMed] [Google Scholar]
24. Reinemer P, Dirr HW, Ladenstein R, Huber R, Lo Bello M, Federici G, Parker MW. J. Mol. Biol. 1992;227:214–226. [PubMed] [Google Scholar]
25. Sinning I, Kleywegt GJ, Cowan SW, Reinemer P, Dirr HW, Huber R, Gilliland GL, Armstrong RN, Ji X, Board PG, et al. J. Mol. Biol. 1993;232:192–212. [PubMed] [Google Scholar]
26. Jencks WP. Catalysis in Chemistry and Enzymology. McGraw-Hill, Inc.; New York, NY: 1969. pp. 585–586. [Google Scholar]
27. Sakurai K, Oobatake M, Goto Y. Protein Sci. 2001;10:2325–2335.[PMC free article] [PubMed] [Google Scholar]
28. Ji X, Tordova M, O'Donnell R, Parsons JF, Hayden JB, Gilliland GL, Zimniak P. Biochemistry. 1997;36:9690–9702. [PubMed] [Google Scholar]
29. Mao L, Wang Y, Lui Y, Hu X. J. Mol. Biol. 2004;336:787–807. [PubMed] [Google Scholar]
30. Adler V, Yin Z, Fuchs SY, Benezra M, Rosario L, Tew KD, Pincus MR, Sardana M, Henderson CJ, Wolf CR, Davis RJ, Ronai Z. EMBO J. 1999;18:1321–1334.[PMC free article] [PubMed] [Google Scholar]
31. Wang T, Arifoglu P, Ronai Z, Tew KD. J. Biol. Chem. 2001;276:20999–21003. [PubMed] [Google Scholar]
32. Elsby R, Kitteringham NR, Goldring CE, Lovatt CA, Chamberlain M, Henderson CJ, Wolf CR, Park BK. J. Biol. Chem. 2003;278:22243–22249. [PubMed] [Google Scholar]
33. Wu Y, Fan Y, Xue B, Luo L, Shen J, Zhang S, Jiang Y, Yin Z. Oncogene. 2006;25:5787–5800. [PubMed] [Google Scholar]
